+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11862 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
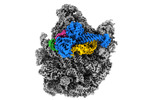
| Title | Structure of the bacterial RQC complex (Decoding State) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RIBOSOME / rqch / rqc / rqc2 / hsp15 / fibronectin-binding protein / NEMF / ribosome-associated quality control / ribosome-associated / 50s / Alanine-tailing | |||||||||
| Function / homology |  Function and homology information Function and homology informationRQC complex / positive regulation of rRNA processing / nucleoid / ribosomal large subunit binding / rescue of stalled ribosome / rRNA processing / large ribosomal subunit / transferase activity / 5S rRNA binding / large ribosomal subunit rRNA binding ...RQC complex / positive regulation of rRNA processing / nucleoid / ribosomal large subunit binding / rescue of stalled ribosome / rRNA processing / large ribosomal subunit / transferase activity / 5S rRNA binding / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / rRNA binding / negative regulation of translation / ribosome / structural constituent of ribosome / translation / ribonucleoprotein complex / response to antibiotic / mRNA binding / DNA binding / RNA binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.99 Å | |||||||||
 Authors Authors | Filbeck S / Pfeffer S | |||||||||
| Funding support |  Germany, Germany,  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2021 Journal: Mol Cell / Year: 2021Title: Mimicry of Canonical Translation Elongation Underlies Alanine Tail Synthesis in RQC. Authors: Sebastian Filbeck / Federico Cerullo / Helge Paternoga / George Tsaprailis / Claudio A P Joazeiro / Stefan Pfeffer /   Abstract: Aborted translation produces large ribosomal subunits obstructed with tRNA-linked nascent chains, which are substrates of ribosome-associated quality control (RQC). Bacterial RqcH, a widely conserved ...Aborted translation produces large ribosomal subunits obstructed with tRNA-linked nascent chains, which are substrates of ribosome-associated quality control (RQC). Bacterial RqcH, a widely conserved RQC factor, senses the obstruction and recruits tRNA to modify nascent-chain C termini with a polyalanine degron. However, how RqcH and its eukaryotic homologs (Rqc2 and NEMF), despite their relatively simple architecture, synthesize such C-terminal tails in the absence of a small ribosomal subunit and mRNA has remained unknown. Here, we present cryoelectron microscopy (cryo-EM) structures of Bacillus subtilis RQC complexes representing different Ala tail synthesis steps. The structures explain how tRNA is selected via anticodon reading during recruitment to the A-site and uncover striking hinge-like movements in RqcH leading tRNA into a hybrid A/P-state associated with peptidyl-transfer. Finally, we provide structural, biochemical, and molecular genetic evidence identifying the Hsp15 homolog (encoded by rqcP) as a novel RQC component that completes the cycle by stabilizing the P-site tRNA conformation. Ala tailing thus follows mechanistic principles surprisingly similar to canonical translation elongation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11862.map.gz emd_11862.map.gz | 7.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11862-v30.xml emd-11862-v30.xml emd-11862.xml emd-11862.xml | 46.5 KB 46.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_11862.png emd_11862.png | 169.2 KB | ||
| Filedesc metadata |  emd-11862.cif.gz emd-11862.cif.gz | 10.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11862 http://ftp.pdbj.org/pub/emdb/structures/EMD-11862 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11862 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11862 | HTTPS FTP |
-Validation report
| Summary document |  emd_11862_validation.pdf.gz emd_11862_validation.pdf.gz | 419 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_11862_full_validation.pdf.gz emd_11862_full_validation.pdf.gz | 418.6 KB | Display | |
| Data in XML |  emd_11862_validation.xml.gz emd_11862_validation.xml.gz | 6.8 KB | Display | |
| Data in CIF |  emd_11862_validation.cif.gz emd_11862_validation.cif.gz | 7.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11862 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11862 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11862 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11862 | HTTPS FTP |
-Related structure data
| Related structure data |  7aqcMC  7aqdC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11862.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11862.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Bacterial RQC complex
+Supramolecule #1: Bacterial RQC complex
+Macromolecule #1: 23S ribosomal RNA
+Macromolecule #2: 5S ribosomal RNA
+Macromolecule #15: Ala-tRNA (P-site)
+Macromolecule #3: 50S ribosomal protein L2
+Macromolecule #4: 50S ribosomal protein L3
+Macromolecule #5: 50S ribosomal protein L4
+Macromolecule #6: 50S ribosomal protein L5
+Macromolecule #7: 50S ribosomal protein L6
+Macromolecule #8: 50S ribosomal protein L11
+Macromolecule #9: 50S ribosomal protein L13
+Macromolecule #10: 50S ribosomal protein L14
+Macromolecule #11: 50S ribosomal protein L15
+Macromolecule #12: 50S ribosomal protein L16
+Macromolecule #13: 50S ribosomal protein L17
+Macromolecule #14: 50S ribosomal protein L18
+Macromolecule #16: 50S ribosomal protein L20
+Macromolecule #17: Rqc2 homolog RqcH
+Macromolecule #18: 50S ribosomal protein L22
+Macromolecule #19: 50S ribosomal protein L24
+Macromolecule #20: 50S ribosomal protein L27
+Macromolecule #21: nascent polyalanine
+Macromolecule #22: 50S ribosomal protein L28
+Macromolecule #23: Uncharacterized protein YabO
+Macromolecule #24: 50S ribosomal protein L30
+Macromolecule #25: 50S ribosomal protein L19
+Macromolecule #26: 50S ribosomal protein L32
+Macromolecule #27: 50S ribosomal protein L33 1
+Macromolecule #28: 50S ribosomal protein L34
+Macromolecule #29: 50S ribosomal protein L35
+Macromolecule #30: 50S ribosomal protein L36
+Macromolecule #31: 50S ribosomal protein L21
+Macromolecule #32: 50S ribosomal protein L23
+Macromolecule #33: 50S ribosomal protein L29
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Output model |  PDB-7aqc: |
 Movie
Movie Controller
Controller