+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11069 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Smooth muscle myosin shutdown state heads region | ||||||||||||
 Map data Map data | Smooth muscle myosin shutdown heads region map | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | myosin / motor / inhibited state / shutdown state / smooth muscle / CONTRACTILE PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationmyosin filament / actomyosin structure organization / myosin II complex / structural constituent of muscle / microfilament motor activity / myofibril / supramolecular fiber organization / ADP binding / actin filament binding / calmodulin binding ...myosin filament / actomyosin structure organization / myosin II complex / structural constituent of muscle / microfilament motor activity / myofibril / supramolecular fiber organization / ADP binding / actin filament binding / calmodulin binding / calcium ion binding / ATP binding / cytosol Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
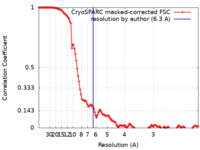
| Method | single particle reconstruction / cryo EM / Resolution: 6.3 Å | ||||||||||||
 Authors Authors | Scarff CA / Carrington G / Casas Mao D / Chalovich JM / Knight PJ / Ranson NA / Peckham M | ||||||||||||
| Funding support |  United Kingdom, 3 items United Kingdom, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nature / Year: 2020 Journal: Nature / Year: 2020Title: Structure of the shutdown state of myosin-2. Authors: Charlotte A Scarff / Glenn Carrington / David Casas-Mao / Joseph M Chalovich / Peter J Knight / Neil A Ranson / Michelle Peckham /   Abstract: Myosin-2 is essential for processes as diverse as cell division and muscle contraction. Dephosphorylation of its regulatory light chain promotes an inactive, 'shutdown' state with the filament- ...Myosin-2 is essential for processes as diverse as cell division and muscle contraction. Dephosphorylation of its regulatory light chain promotes an inactive, 'shutdown' state with the filament-forming tail folded onto the two heads, which prevents filament formation and inactivates the motors. The mechanism by which this happens is unclear. Here we report a cryo-electron microscopy structure of shutdown smooth muscle myosin with a resolution of 6 Å in the head region. A pseudo-atomic model, obtained by flexible fitting of crystal structures into the density and molecular dynamics simulations, describes interaction interfaces at the atomic level. The N-terminal extension of one regulatory light chain interacts with the tail, and the other with the partner head, revealing how the regulatory light chains stabilize the shutdown state in different ways and how their phosphorylation would allow myosin activation. Additional interactions between the three segments of the coiled coil, the motor domains and the light chains stabilize the shutdown molecule. The structure of the lever in each head is competent to generate force upon activation. This shutdown structure is relevant to all isoforms of myosin-2 and provides a framework for understanding their disease-causing mutations. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11069.map.gz emd_11069.map.gz | 6.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11069-v30.xml emd-11069-v30.xml emd-11069.xml emd-11069.xml | 20.8 KB 20.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11069_fsc.xml emd_11069_fsc.xml | 13.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_11069.png emd_11069.png | 31.3 KB | ||
| Filedesc metadata |  emd-11069.cif.gz emd-11069.cif.gz | 7.1 KB | ||
| Others |  emd_11069_half_map_1.map.gz emd_11069_half_map_1.map.gz emd_11069_half_map_2.map.gz emd_11069_half_map_2.map.gz | 154.6 MB 154.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11069 http://ftp.pdbj.org/pub/emdb/structures/EMD-11069 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11069 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11069 | HTTPS FTP |
-Related structure data
| Related structure data |  6z47MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11069.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11069.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Smooth muscle myosin shutdown heads region map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: Smooth muscle myosin shutdown heads region half map A
| File | emd_11069_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Smooth muscle myosin shutdown heads region half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Smooth muscle myosin shutdown heads region half map B
| File | emd_11069_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Smooth muscle myosin shutdown heads region half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Smooth muscle myosin in the shutdown state
| Entire | Name: Smooth muscle myosin in the shutdown state |
|---|---|
| Components |
|
-Supramolecule #1: Smooth muscle myosin in the shutdown state
| Supramolecule | Name: Smooth muscle myosin in the shutdown state / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Myosin heavy chain 11
| Macromolecule | Name: Myosin heavy chain 11 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 229.14825 KDa |
| Sequence | String: MSQKPLSDDE KFLFVDKNFV NNPLAQADWS AKKLVWVPSE KHGFEAASIK EEKGDEVTVE LQENGKKVTL SKDDIQKMNP PKFSKVEDM AELTCLNEAS VLHNLRERYF SGLIYTYSGL FCVVVNPYKQ LPIYSEKIID MYKGKKRHEM PPHIYAIADT A YRSMLQDR ...String: MSQKPLSDDE KFLFVDKNFV NNPLAQADWS AKKLVWVPSE KHGFEAASIK EEKGDEVTVE LQENGKKVTL SKDDIQKMNP PKFSKVEDM AELTCLNEAS VLHNLRERYF SGLIYTYSGL FCVVVNPYKQ LPIYSEKIID MYKGKKRHEM PPHIYAIADT A YRSMLQDR EDQSILCTGE SGAGKTENTK KVIQYLAVVA SSHKGKKDTS ITQGPSFSYG ELEKQLLQAN PILEAFGNAK TV KNDNSSR FGKFIRINFD VTGYIVGANI ETYLLEKSRA IRQAKDERTF HIFYYLIAGA SEQMRNDLLL EGFNNYTFLS NGH VPIPAQ QDDEMFQETL EAMRIMGFTE EEQTSILRVV SSVLQLGNIV FKKERNTDQA SMPDNTAAQK VCHLMGINVT DFTR SILTP RIKVGRDVVQ KAQTKEQADF AIEALAKAKF ERLFRWILTR VNKALDKTKR QGASFLGILD IAGFEIFEIN SFEQL CINY TNEKLQQLFN HTMFILEQEE YQREGIEWNF IDFGLDLQPC IELIERPTNP PGVLALLDEE CWFPKATDTS FVEKLI QEQ GNHPKFQKSK QLKDKTEFCI LHYAGKVSYN ASAWLTKNMD PLNDNVTSLL NQSSDKFVAD LWKDVDRIVG LDQMAKM TE SSLPSSSKTK KGMFRTVGQL YKEQLTKLMT TLRNTNPNFV RCIIPNHEKR AGKLDAHLVL EQLRCNGVLE GIRICRQG F PNRIVFQEFR QRYEILAANA IPKGFMDGKQ ACILMIKALE LDPNLYRIGQ SKIFFRTGVL AHLEEERDLK ITDVIIAFQ AQCRGYLARK AFAKRQQQLT AMKVIQRNCA AYLKLRNWQW WRLFTKVKPL LQVTRQEEEM QAKDEELQRT KERQQKAEAE LKELEQKHT QLCEEKNLLQ EKLQAETELY AEAEEMRVRL AAKKQELEEI LHEMEARIEE EEERSQQLQA EKKKMQQQML D LEEQLEEE EAARQKLQLE KVTADGKIKK MEDDILIMED QNNKLTKERK LLEERVSDLT TNLAEEEEKA KNLTKLKNKH ES MISELEV RLKKEEKTRQ ELEKTKRKLE GESSDLHEQI AELQAQIAEL KAQLAKKEEE LQAALARLED ETSQKNNALK KIR ELESHI SDLQEDLESE KAARNKAEKQ KRDLGEELEA LKTELEDTLD TTATQQELRA KREQEVTVLK RALEEETRTH EAQV QEMRQ KHTQAVEELT EQLEQFKRAK ANLDKTKQTL EKDNADLANE VRSLSQAKQD VEHKKKKLEV QLQDLQSKYT DGERV RTEL NEKVHKLQIE VENVTSLLNE AESKNIKLTK DVATLGSQLQ DTQELLQEET RQKLNVTTKL RQLEDDKNSL QEQLDE EVE AKQNLERHIS TLTIQLSDSK KKLQEFTATI ETMEEGKKKF QREIESLTQQ FEEKAASYDK LEKTKNRLQQ ELDDLVV DL DNQRQLVSNL EKKQKKFDQM LAEEKNISSK YADERDRAEA EAREKETKAL SLARALEEAL EAKEELERTN KMLKAEME D LVSSKDDVGK NVHELEKSKR TLEQQVEEMK TQLEELEDEL QAAEDAKLRL EVNMQAMKSQ FERDLQARDE QNEEKRRQL LKQLHEHETE LEDERKQRAL AAAAKKKLEV DVKDLESQVD SVNKAREEAI KQLRKLQAQM KDYQRDLDDA RAAREEIFAT ARENEKKAK NLEAELIQLQ EDLAAAERAR KQADLEKEEM AEELASATSG RTSLQDDKRR LEARIAQLEE ELDEEHSNIE A MSDRMRKA VQQAEQLNNE LATERATAQK NENARQQLER QNKELRSKLQ EMEGAVKSKF KSTIAALEAK IASLEEQLEQ EA REKQAAA KTLRQKDKKL KDALLQVEDE KKQAEQYKDQ AEKGNLRLKQ LKRQLEEAEE ESQRINANRR KLQRELDEAT ESN DALGRE VAALKSKLRR GNEPVSFAPP RRSGGRRVIE NATDGGEQEI DGRDGDLNGK ASE UniProtKB: Myosin heavy chain 11 |
-Macromolecule #2: Myosin light chain smooth muscle isoform
| Macromolecule | Name: Myosin light chain smooth muscle isoform / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 16.989145 KDa |
| Sequence | String: MCDFSEEQTA EFKEAFQLFD RTGDGKILYS QCGDVMRALG QNPTNAEVMK VLGNPKSDEM NLKTLNFEQF LPMMQTIAKN KDQGCFEDY VEGLRVFDKE GNGTVMGAEI RHVLVTLGEK MTEEEVEQLV AGHEDSNGCI NYEELVRMVL SG UniProtKB: Myosin light chain smooth muscle isoform |
-Macromolecule #3: Myosin light chain 9
| Macromolecule | Name: Myosin light chain 9 / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 19.872244 KDa |
| Sequence | String: MSSKRAKAKT TKKRPQRATS NVFAMFDQSQ IQEFKEAFNM IDQNRDGFID KEDLHDMLAS MGKNPTDEYL EGMMSEAPGP INFTMFLTM FGEKLNGTDP EDVIRNAFAC FDEEASGFIH EDHLRELLTT MGDRFTDEEV DEMYREAPID KKGNFNYVEF T RILKHGAK DKDD UniProtKB: Myosin light chain 9 |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 4 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 5 / Number of copies: 2 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #6: PHOSPHATE ION
| Macromolecule | Name: PHOSPHATE ION / type: ligand / ID: 6 / Number of copies: 2 / Formula: PO4 |
|---|---|
| Molecular weight | Theoretical: 94.971 Da |
| Chemical component information |  ChemComp-PO4: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.45 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.2 Component:
| ||||||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AMYLAMINE / Details: GloCube | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 281 K / Instrument: FEI VITROBOT MARK IV / Details: Blot force 6, blot time 3 seconds. | ||||||||||||
| Details | Smooth muscle myosin from turkey gizzard, stored under liquid nitrogen, was thawed quickly. MgATP was added before dilution into the final buffer conditions stated below |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)