+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10794 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Photorhabdus luminescens TcdA1 in complex with BSA-Lewis X | |||||||||
 Map data Map data | The main map of Pl-TcdA1 with BSA-LewisX after postprocessing | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Photorhabdus luminescens (bacteria) / Photorhabdus luminescens (bacteria) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 5.0 Å | |||||||||
 Authors Authors | Roderer D / Broecker F / Sitsel O / Kaplonek P / Leidreiter F / Seeberger PH / Raunser S | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: Glycan-dependent cell adhesion mechanism of Tc toxins. Authors: Daniel Roderer / Felix Bröcker / Oleg Sitsel / Paulina Kaplonek / Franziska Leidreiter / Peter H Seeberger / Stefan Raunser /   Abstract: Toxin complex (Tc) toxins are virulence factors of pathogenic bacteria. Tcs are composed of three subunits: TcA, TcB and TcC. TcA facilitates receptor-toxin interaction and membrane permeation, TcB ...Toxin complex (Tc) toxins are virulence factors of pathogenic bacteria. Tcs are composed of three subunits: TcA, TcB and TcC. TcA facilitates receptor-toxin interaction and membrane permeation, TcB and TcC form a toxin-encapsulating cocoon. While the mechanisms of holotoxin assembly and pore formation have been described, little is known about receptor binding of TcAs. Here, we identify heparins/heparan sulfates and Lewis antigens as receptors for different TcAs from insect and human pathogens. Glycan array screening reveals that all tested TcAs bind negatively charged heparins. Cryo-EM structures of Morganella morganii TcdA4 and Xenorhabdus nematophila XptA1 reveal that heparins/heparan sulfates unexpectedly bind to different regions of the shell domain, including receptor-binding domains. In addition, Photorhabdus luminescens TcdA1 binds to Lewis antigens with micromolar affinity. Here, the glycan interacts with the receptor-binding domain D of the toxin. Our results suggest a glycan dependent association mechanism of Tc toxins on the host cell surface. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10794.map.gz emd_10794.map.gz | 15.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10794-v30.xml emd-10794-v30.xml emd-10794.xml emd-10794.xml | 16.4 KB 16.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_10794.png emd_10794.png | 164.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10794 http://ftp.pdbj.org/pub/emdb/structures/EMD-10794 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10794 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10794 | HTTPS FTP |
-Validation report
| Summary document |  emd_10794_validation.pdf.gz emd_10794_validation.pdf.gz | 217.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_10794_full_validation.pdf.gz emd_10794_full_validation.pdf.gz | 217.1 KB | Display | |
| Data in XML |  emd_10794_validation.xml.gz emd_10794_validation.xml.gz | 7.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10794 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10794 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10794 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10794 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10794.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10794.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The main map of Pl-TcdA1 with BSA-LewisX after postprocessing | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
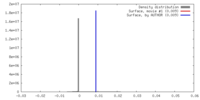
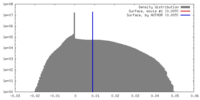
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Photorhabdus luminescens TcdA1 pentamer with one BSA-Lewis X, cro...
| Entire | Name: Photorhabdus luminescens TcdA1 pentamer with one BSA-Lewis X, crosslinked with glutaraldehyde |
|---|---|
| Components |
|
-Supramolecule #1: Photorhabdus luminescens TcdA1 pentamer with one BSA-Lewis X, cro...
| Supramolecule | Name: Photorhabdus luminescens TcdA1 pentamer with one BSA-Lewis X, crosslinked with glutaraldehyde type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Molecular weight | Theoretical: 1.5 MDa |
-Supramolecule #2: Photorhabdus luminescens toxin complex subunit TcdA1, Homopentamer
| Supramolecule | Name: Photorhabdus luminescens toxin complex subunit TcdA1, Homopentamer type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Photorhabdus luminescens (bacteria) Photorhabdus luminescens (bacteria) |
| Recombinant expression | Organism:  |
-Supramolecule #3: BSA with Lewis X glycans crosslinked to surface lysines
| Supramolecule | Name: BSA with Lewis X glycans crosslinked to surface lysines type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Photorhabdus luminescens toxin complex subunit TcdA1, Homopentamer
| Macromolecule | Name: Photorhabdus luminescens toxin complex subunit TcdA1, Homopentamer type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Photorhabdus luminescens (bacteria) Photorhabdus luminescens (bacteria) |
| Recombinant expression | Organism:  |
| Sequence | String: MNESVKEIPD VLKSQCGFNC LTDISHSSFN EFRQQVSEHL SWSETHDLYH DAQQAQKDNR LYEARILKR ANPQLQNAVH LAILAPNAEL IGYNNQFSGR ASQYVAPGTV SSMFSPAAYL T ELYREARN LHASDSVYYL DTRRPDLKSM ALSQQNMDIE LSTLSLSNEL ...String: MNESVKEIPD VLKSQCGFNC LTDISHSSFN EFRQQVSEHL SWSETHDLYH DAQQAQKDNR LYEARILKR ANPQLQNAVH LAILAPNAEL IGYNNQFSGR ASQYVAPGTV SSMFSPAAYL T ELYREARN LHASDSVYYL DTRRPDLKSM ALSQQNMDIE LSTLSLSNEL LLESIKTESK LE NYTKVME MLSTFRPSGA TPYHDAYENV REVIQLQDPG LEQLNASPAI AGLMHQASLL GIN ASISPE LFNILTEEIT EGNAEELYKK NFGNIEPASL AMPEYLKRYY NLSDEELSQF IGKA SNFGQ QEYSNNQLIT PVVNSSDGTV KVYRITREYT TNAYQMDVEL FPFGGENYRL DYKFK NFYN ASYLSIKLND KRELVRTEGA PQVNIEYSAN ITLNTADISQ PFEIGLTRVL PSGSWA YAA AKFTVEEYNQ YSFLLKLNKA IRLSRATELS PTILEGIVRS VNLQLDINTD VLGKVFL TK YYMQRYAIHA ETALILCNAP ISQRSYDNQP SQFDRLFNTP LLNGQYFSTG DEEIDLNS G STGDWRKTIL KRAFNIDDVS LFRLLKITDH DNKDGKIKNN LKNLSNLYIG KLLADIHQL TIDELDLLLI AVGEGKTNLS AISDKQLATL IRKLNTITSW LHTQKWSVFQ LFIMTSTSYN KTLTPEIKN LLDTVYHGLQ GFDKDKADLL HVMAPYIAAT LQLSSENVAH SVLLWADKLQ P GDGAMTAE KFWDWLNTKY TPGSSEAVET QEHIVQYCQA LAQLEMVYHS TGINENAFRL FV TKPEMFG AATGAAPAHD ALSLIMLTRF ADWVNALGEK ASSVLAAFEA NSLTAEQLAD AMN LDANLL LQASIQAQNH QHLPPVTPEN AFSCWTSINT ILQWVNVAQQ LNVAPQGVSA LVGL DYIQS MKETPTYAQW ENAAGVLTAG LNSQQANTLH AFLDESRSAA LSTYYIRQVA KAAAA IKSR DDLYQYLLID NQVSAAIKTT RIAEAIASIQ LYVNRALENV EENANSGVIS RQFFID WDK YNKRYSTWAG VSQLVYYPEN YIDPTMRIGQ TKMMDALLQS VSQSQLNADT VEDAFMS YL TSFEQVANLK VISAYHDNIN NDQGLTYFIG LSETDAGEYY WRSVDHSKFN DGKFAANA W SEWHKIDCPI NPYKSTIRPV IYKSRLYLLW LEQKEITKQT GNSKDGYQTE TDYRYELKL AHIRYDGTWN TPITFDVNKK ISELKLEKNR APGLYCAGYQ GEDTLLVMFY NQQDTLDSYK NASMQGLYI FADMASKDMT PEQSNVYRDN SYQQFDTNNV RRVNNRYAED YEIPSSVSSR K DYGWGDYY LSMVYNGDIP TINYKAASSD LKIYISPKLR IIHNGYEGQK RNQCNLMNKY GK LGDKFIV YTSLGVNPNN SSNKLMFYPV YQYSGNTSGL NQGRLLFHRD TTYPSKVEAW IPG AKRSLT NQNAAIGDDY ATDSLNKPDD LKQYIFMTDS KGTATDVSGP VEINTAISPA KVQI IVKAG GKEQTFTADK DVSIQPSPSF DEMNYQFNAL EIDGSGLNFI NNSASIDVTF TAFAE DGRK LGYESFSIPV TLKVSTDNAL TLHHNENGAQ YMQWQSYRTR LNTLFARQLV ARATTG IDT ILSMETQNIQ EPQLGKGFYA TFVIPPYNLS THGDERWFKL YIKHVVDNNS HIIYSGQ LT DTNINITLFI PLDDVPLNQD YHAKVYMTFK KSPSDGTWWG PHFVRDDKGI VTINPKSI L THFESVNVLN NISSEPMDFS GANSLYFWEL FYYTPMLVAQ RLLHEQNFDE ANRWLKYVW SPSGYIVHGQ IQNYQWNVRP LLEDTSWNSD PLDSVDPDAV AQHDPMHYKV STFMRTLDLL IARGDHAYR QLERDTLNEA KMWYMQALHL LGDKPYLPLS TTWSDPRLDR AADITTQNAH D SAIVALRQ NIPTPAPLSL RSANTLTDLF LPQINEVMMN YWQTLAQRVY NLRHNLSIDG QP LYLPIYA TPADPKALLS AAVATSQGGG KLPESFMSLW RFPHMLENAR GMVSQLTQFG STL QNIIER QDAEALNALL QNQAAELILT NLSIQDKTIE ELDAEKTVLE KSKAGAQSRF DSYG KLYDE NINAGENQAM TLRASAAGLT TAVQASRLAG AAADLVPNIF GFAGGGSRWG AIAEA TGYV MEFSANVMNT EADKISQSET YRRRRQEWEI QRNNAEAELK QIDAQLKSLA VRREAA VLQ KTSLKTQQEQ TQSQLAFLQR KFSNQALYNW LRGRLAAIYF QFYDLAVARC LMAEQAY RW ELNDDSARFI KPGAWQGTYA GLLAGETLML SLAQMEDAHL KRDKRALEVE RTVSLAEV Y AGLPKDNGPF SLAQEIDKLV SQGSGSAGSG NNNLAFGAGT DTKTSLQASV SFADLKIRE DYPASLGKIR RIKQISVTLP ALLGPYQDVQ AILSYGDKAG LANGCEALAV SHGMNDSGQF QLDFNDGKF LPFEGIAIDQ GTLTLSFPNA SMPEKGKQAT MLKTLNDIIL HIRYTIK |
-Macromolecule #2: BSA with Lewis X glycans crosslinked to surface lysines
| Macromolecule | Name: BSA with Lewis X glycans crosslinked to surface lysines type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MKWVTFISLL LLFSSAYSRG VFRRDTHKSE IAHRFKDLGE EHFKGLVLIA FSQYLQQCPF DEHVKLVNE LTEFAKTCVA DESHAGCEKS LHTLFGDELC KVASLRETYG DMADCCEKQE P ERNECFLS HKDDSPDLPK LKPDPNTLCD EFKADEKKFW GKYLYEIARR ...String: MKWVTFISLL LLFSSAYSRG VFRRDTHKSE IAHRFKDLGE EHFKGLVLIA FSQYLQQCPF DEHVKLVNE LTEFAKTCVA DESHAGCEKS LHTLFGDELC KVASLRETYG DMADCCEKQE P ERNECFLS HKDDSPDLPK LKPDPNTLCD EFKADEKKFW GKYLYEIARR HPYFYAPELL YY ANKYNGV FQECCQAEDK GACLLPKIET MREKVLASSA RQRLRCASIQ KFGERALKAW SVA RLSQKF PKAEFVEVTK LVTDLTKVHK ECCHGDLLEC ADDRADLAKY ICDNQDTISS KLKE CCDKP LLEKSHCIAE VEKDAIPENL PPLTADFAED KDVCKNYQEA KDAFLGSFLY EYSRR HPEY AVSVLLRLAK EYEATLEECC AKDDPHACYS TVFDKLKHLV DEPQNLIKQN CDQFEK LGE YGFQNALIVR YTRKVPQVST PTLVEVSRSL GKVGTRCCTK PESERMPCTE DYLSLIL NR LCVLHEKTPV SEKVTKCCTE SLVNRRPCFS ALTPDETYVP KAFDEKLFTF HADICTLP D TEKQIKKQTA LVELLKHKPK ATEEQLKTVM ENFVAFVDKC CAADDKEACF AVEGPKLVV STQTALA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.06 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2.0 nm / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 293 K / Instrument: GATAN CRYOPLUNGE 3 |
| Details | The sample was crosslinked with glutaraldehyde and subsequently purified by SEC |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Number real images: 4992 / Average electron dose: 100.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)