+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10361 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human Influenza B polymerase recycling complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Influenza / polymerase / viral transcription / RNA / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcap snatching / viral transcription / symbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity / host cell mitochondrion / 7-methylguanosine mRNA capping / virion component / endonuclease activity / Hydrolases; Acting on ester bonds / host cell cytoplasm / symbiont-mediated suppression of host gene expression ...cap snatching / viral transcription / symbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity / host cell mitochondrion / 7-methylguanosine mRNA capping / virion component / endonuclease activity / Hydrolases; Acting on ester bonds / host cell cytoplasm / symbiont-mediated suppression of host gene expression / viral translational frameshifting / RNA-directed RNA polymerase / nucleotide binding / viral RNA genome replication / hydrolase activity / RNA-directed RNA polymerase activity / DNA-templated transcription / host cell nucleus / RNA binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Influenza B virus Influenza B virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.18 Å | |||||||||
 Authors Authors | Wandzik JM / Kouba T | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2020 Journal: Cell / Year: 2020Title: A Structure-Based Model for the Complete Transcription Cycle of Influenza Polymerase. Authors: Joanna M Wandzik / Tomas Kouba / Manikandan Karuppasamy / Alexander Pflug / Petra Drncova / Jan Provaznik / Nayara Azevedo / Stephen Cusack /   Abstract: Influenza polymerase uses unique mechanisms to synthesize capped and polyadenylated mRNAs from the genomic viral RNA (vRNA) template, which is packaged inside ribonucleoprotein particles (vRNPs). ...Influenza polymerase uses unique mechanisms to synthesize capped and polyadenylated mRNAs from the genomic viral RNA (vRNA) template, which is packaged inside ribonucleoprotein particles (vRNPs). Here, we visualize by cryoelectron microscopy the conformational dynamics of the polymerase during the complete transcription cycle from pre-initiation to termination, focusing on the template trajectory. After exiting the active site cavity, the template 3' extremity rebinds into a specific site on the polymerase surface. Here, it remains sequestered during all subsequent transcription steps, forcing the template to loop out as it further translocates. At termination, the strained connection between the bound template 5' end and the active site results in polyadenylation by stuttering at uridine 17. Upon product dissociation, further conformational changes release the trapped template, allowing recycling back into the pre-initiation state. Influenza polymerase thus performs transcription while tightly binding to and protecting both template ends, allowing efficient production of multiple mRNAs from a single vRNP. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10361.map.gz emd_10361.map.gz | 19.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10361-v30.xml emd-10361-v30.xml emd-10361.xml emd-10361.xml | 25 KB 25 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10361_fsc.xml emd_10361_fsc.xml | 6.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_10361.png emd_10361.png | 89.6 KB | ||
| Filedesc metadata |  emd-10361.cif.gz emd-10361.cif.gz | 7.8 KB | ||
| Others |  emd_10361_additional.map.gz emd_10361_additional.map.gz emd_10361_half_map_1.map.gz emd_10361_half_map_1.map.gz emd_10361_half_map_2.map.gz emd_10361_half_map_2.map.gz | 9.9 MB 15.9 MB 15.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10361 http://ftp.pdbj.org/pub/emdb/structures/EMD-10361 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10361 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10361 | HTTPS FTP |
-Related structure data
| Related structure data |  6t0wMC  6szuC  6szvC  6t0nC  6t0rC  6t0sC  6t0uC  6t0vC  6t2cC  6tu5C  6tw1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10361.map.gz / Format: CCP4 / Size: 20.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10361.map.gz / Format: CCP4 / Size: 20.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0506 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
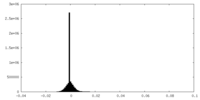
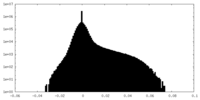
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: LocScale map
| File | emd_10361_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | LocScale map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_10361_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_10361_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human Influenza B polymerase recycling complex
| Entire | Name: Human Influenza B polymerase recycling complex |
|---|---|
| Components |
|
-Supramolecule #1: Human Influenza B polymerase recycling complex
| Supramolecule | Name: Human Influenza B polymerase recycling complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: Polymerase
| Supramolecule | Name: Polymerase / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Influenza B virus Influenza B virus |
-Supramolecule #3: Nucleic acid
| Supramolecule | Name: Nucleic acid / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #4 |
|---|---|
| Source (natural) | Organism:  Influenza B virus Influenza B virus |
-Macromolecule #1: Polymerase acidic protein
| Macromolecule | Name: Polymerase acidic protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: Hydrolases; Acting on ester bonds |
|---|---|
| Source (natural) | Organism:  Influenza B virus Influenza B virus |
| Molecular weight | Theoretical: 85.822781 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: GSHHHHHHHH GSGSMDTFIT RNFQTTIIQK AKNTMAEFSE DPELQPAMLF NICVHLEVCY VISDMNFLDE EGKAYTALEG QGKEQNLRP QYEVIEGMPR TIAWMVQRSL AQEHGIETPK YLADLFDYKT KRFIEVGITK GLADDYFWKK KEKLGNSMEL M IFSYNQDY ...String: GSHHHHHHHH GSGSMDTFIT RNFQTTIIQK AKNTMAEFSE DPELQPAMLF NICVHLEVCY VISDMNFLDE EGKAYTALEG QGKEQNLRP QYEVIEGMPR TIAWMVQRSL AQEHGIETPK YLADLFDYKT KRFIEVGITK GLADDYFWKK KEKLGNSMEL M IFSYNQDY SLSNESSLDE EGKGRVLSRL TELQAELSLK NLWQVLIGEE DVEKGIDFKL GQTISRLRDI SVPAGFSNFE GM RSYIDNI DPKGAIERNL ARMSPLVSVT PKKLTWEDLR PIGPHIYNHE LPEVPYNAFL LMSDELGLAN MTEGKSKKPK TLA KECLEK YSTLRDQTDP ILIMKSEKAN ENFLWKLWRD CVNTISNEEM SNELQKTNYA KWATGDGLTY QKIMKEVAID DETM CQEEP KIPNKCRVAA WVQTEMNLLS TLTSKRALDL PEIGPDVAPV EHVGSERRKY FVNEINYCKA STVMMKYVLF HTSLL NESN ASMGKYKVIP ITNRVVNEKG ESFDMLYGLA VKGQSHLRGD TDVVTVVTFE FSSTDPRVDS GKWPKYTVFR IGSLFV SGR EKSVYLYCRV NGTNKIQMKW GMEARRCLLQ SMQQMEAIVE QESSIQGYDM TKACFKGDRV NSPKTFSIGT QEGKLVK GS FGKALRVIFT KCLMHYVFGN AQLEGFSAES RRLLLLIQAL KDRKGPWVFD LEGMYSGIEE CISNNPWVIQ SAYWFNEW L GFEKEGSKVL ESVDEIMDEG SGSGENLYFQ UniProtKB: Polymerase acidic protein |
-Macromolecule #2: RNA-directed RNA polymerase catalytic subunit
| Macromolecule | Name: RNA-directed RNA polymerase catalytic subunit / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: RNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  Influenza B virus Influenza B virus |
| Molecular weight | Theoretical: 86.207016 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: GSGSGSGSGM NINPYFLFID VPIQAAISTT FPYTGVPPYS HGTGTGYTID TVIRTHEYSN KGKQYISDVT GCTMVDPTNG PLPEDNEPS AYAQLDCVLE ALDRMDEEHP GLFQAASQNA METLMVTTVD KLTQGRQTFD WTVCRNQPAA TALNTTITSF R LNDLNGAD ...String: GSGSGSGSGM NINPYFLFID VPIQAAISTT FPYTGVPPYS HGTGTGYTID TVIRTHEYSN KGKQYISDVT GCTMVDPTNG PLPEDNEPS AYAQLDCVLE ALDRMDEEHP GLFQAASQNA METLMVTTVD KLTQGRQTFD WTVCRNQPAA TALNTTITSF R LNDLNGAD KGGLIPFCQD IIDSLDRPEM TFFSVKNIKK KLPAKNRKGF LIKRIPMKVK DKITKVEYIK RALSLNTMTK DA ERGKLKR RAIATAGIQI RGFVLVVENL AKNICENLEQ SGLPVGGNEK KAKLSNAVAK MLSNCPPGGI SMTVTGDNTK WNE CLNPRI FLAMTERITR DSPIWFRDFC SIAPVLFSNK IARLGKGFMI TSKTKRLKAQ IPCPDLFSIP LERYNEETRA KLKK LKPFF NEEGTASLSP GMMMGMFNML STVLGVAALG IKNIGNKEYL WDGLQSSDDF ALFVNAKDEE TCMEGINDFY RTCKL LGIN MSKKKSYCNE TGMFEFTSMF YRDGFVSNFA MELPSFGVAG VNESADMAIG MTIIKNNMIN NGMGPATAQT AIQLFI ADY RYTYKCHRGD SKVEGKRMKI IKELWENTKG RDGLLVADGG PNIYNLRNLH IPEIVLKYNL MDPEYKGRLL HPQNPFV GH LSIEGIKEAD ITPAHGPVKK MDYDAVSGTH SWRTKRNRSI LNTDQRNMIL EEQCYAKCCN LFEACFNSAS YRKPVGQH S MLEAMAHRLR MDARLDYESG RMSKDDFEKA MAHLGEIGYI GSGSGENLYF Q UniProtKB: RNA-directed RNA polymerase catalytic subunit |
-Macromolecule #3: Polymerase PB2
| Macromolecule | Name: Polymerase PB2 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Influenza B virus Influenza B virus |
| Molecular weight | Theoretical: 90.971281 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: GSGSGSGSGM TLAKIELLKQ LLRDNEAKTV LKQTTVDQYN IIRKFNTSRI EKNPSLRMKW AMCSNFPLAL TKGDMANRIP LEYKGIQLK TNAEDIGTKG QMCSIAAVTW WNTYGPIGDT EGFERVYESF FLRKMRLDNA TWGRITFGPV ERVRKRVLLN P LTKEMPPD ...String: GSGSGSGSGM TLAKIELLKQ LLRDNEAKTV LKQTTVDQYN IIRKFNTSRI EKNPSLRMKW AMCSNFPLAL TKGDMANRIP LEYKGIQLK TNAEDIGTKG QMCSIAAVTW WNTYGPIGDT EGFERVYESF FLRKMRLDNA TWGRITFGPV ERVRKRVLLN P LTKEMPPD EASNVIMEIL FPKEAGIPRE STWIHRELIK EKREKLKGTM ITPIVLAYML ERELVARRRF LPVAGATSAE FI EMLHCLQ GENWRQIYHP GGNKLTESRS QSMIVACRKI IRRSIVASNP LELAVEIANK TVIDTEPLKS CLAAIDGGDV ACD IIRAAL GLKIRQRQRF GRLELKRISG RGFKNDEEIL IGNGTIQKIG IWDGEEEFHV RCGECRGILK KSKMKLEKLL INSA KKEDM RDLIILCMVF SQDTRMFQGV RGEINFLNRA GQLLSPMYQL QRYFLNRSND LFDQWGYEES PKASELHGIN ESMNA SDYT LKGVVVTRNV IDDFSSTETE KVSITKNLSL IKRTGEVIMG ANDVSELESQ AQLMITYDTP KMWEMGTTKE LVQNTY QWV LKNLVTLKAQ FLLGKEDMFQ WDAFEAFESI IPQKMAGQYS GFARAVLKQM RDQEVMKTDQ FIKLLPFCFS PPKLRSN GE PYQFLKLVLK GGGENFIEVR KGSPLFSYNP QTEVLTICGR MMSLKGKIED EERNRSMGNA VLAGFLVSGK YDPDLGDF K TIEELEKLKP GEKANILLYQ GKPVKVVKRK RYSALSNDIS QGIKRQRMTV ESMGWALSGW SHPQFEKGRS GGENLYFQ UniProtKB: Polymerase basic protein 2 |
-Macromolecule #4: vRNA
| Macromolecule | Name: vRNA / type: rna / ID: 4 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Influenza B virus Influenza B virus |
| Molecular weight | Theoretical: 10.838431 KDa |
| Sequence | String: AGUAGUAACA AGAGGUAUUA CCUCUGCUUC UGCU |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 35.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)