+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10036 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Photorhabdus luminescens TcdA4 | |||||||||
 Map data Map data | Cryo-EM density map of Photorhabdus luminescens TcdA4 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Toxin / membrane permeation / translocation / complex | |||||||||
| Function / homology |  Function and homology information Function and homology information: / TcdA1, receptor binding domain / TcA receptor binding domain / TcA receptor binding domain / ABC toxin, N-terminal domain / ABC toxin N-terminal region / Insecticidal toxin complex/plasmid virulence protein / Tc toxin complex TcA, C-terminal TcB-binding domain / Neuraminidase-like domain / Salmonella virulence plasmid 28.1kDa A protein ...: / TcdA1, receptor binding domain / TcA receptor binding domain / TcA receptor binding domain / ABC toxin, N-terminal domain / ABC toxin N-terminal region / Insecticidal toxin complex/plasmid virulence protein / Tc toxin complex TcA, C-terminal TcB-binding domain / Neuraminidase-like domain / Salmonella virulence plasmid 28.1kDa A protein / Tc toxin complex TcA C-terminal TcB-binding domain / Neuraminidase-like domain Similarity search - Domain/homology | |||||||||
| Biological species |  Photorhabdus luminescens (bacteria) Photorhabdus luminescens (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||
 Authors Authors | Roderer D / Leidreiter F | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2019 Journal: Sci Adv / Year: 2019Title: Common architecture of Tc toxins from human and insect pathogenic bacteria. Authors: F Leidreiter / D Roderer / D Meusch / C Gatsogiannis / R Benz / S Raunser /  Abstract: Tc toxins use a syringe-like mechanism to penetrate the membrane and translocate toxic enzymes into the host cytosol. They are composed of three components: TcA, TcB, and TcC. Low-resolution ...Tc toxins use a syringe-like mechanism to penetrate the membrane and translocate toxic enzymes into the host cytosol. They are composed of three components: TcA, TcB, and TcC. Low-resolution structures of TcAs from different bacteria suggest a considerable difference in their architecture and possibly in their mechanism of action. Here, we present high-resolution structures of five TcAs from insect and human pathogens, which show a similar overall composition and domain organization. Essential structural features, including a trefoil protein knot, are present in all TcAs, suggesting a common mechanism of action. All TcAs form functional pores and can be combined with TcB-TcC subunits from other species to form active chimeric holotoxins. We identified a conserved ionic pair that stabilizes the shell, likely operating as a strong latch that only springs open after destabilization of other regions. Our results provide new insights into the architecture and mechanism of the Tc toxin family. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10036.map.gz emd_10036.map.gz | 18.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10036-v30.xml emd-10036-v30.xml emd-10036.xml emd-10036.xml | 12.2 KB 12.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_10036.png emd_10036.png | 70.8 KB | ||
| Filedesc metadata |  emd-10036.cif.gz emd-10036.cif.gz | 6.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10036 http://ftp.pdbj.org/pub/emdb/structures/EMD-10036 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10036 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10036 | HTTPS FTP |
-Related structure data
| Related structure data |  6rwaMC  6rw6C  6rw8C  6rw9C  6rwbC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10036.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10036.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM density map of Photorhabdus luminescens TcdA4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.12 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
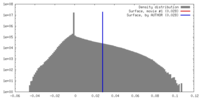
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : P. luminescens TcdA4 pentamer
| Entire | Name: P. luminescens TcdA4 pentamer |
|---|---|
| Components |
|
-Supramolecule #1: P. luminescens TcdA4 pentamer
| Supramolecule | Name: P. luminescens TcdA4 pentamer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Photorhabdus luminescens (bacteria) Photorhabdus luminescens (bacteria) |
| Molecular weight | Theoretical: 1.4 MDa |
-Macromolecule #1: TcdA4
| Macromolecule | Name: TcdA4 / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Photorhabdus luminescens (bacteria) Photorhabdus luminescens (bacteria) |
| Molecular weight | Theoretical: 270.69525 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNSYVKEIPD VLQSQYGINC LTDICHYSFN EFRQQVSDHL SWSETNRLYR DAQQEQKENQ LYEARILKRA NPQLQNAVHL GITLPHAEL RGYNSEFGGR ASQYVAPGSV SSMFSPAAYL TELYREARNL HASDSVYHLD ERRPDLQSMT LSQQNMDTEL S TLSLSNEI ...String: MNSYVKEIPD VLQSQYGINC LTDICHYSFN EFRQQVSDHL SWSETNRLYR DAQQEQKENQ LYEARILKRA NPQLQNAVHL GITLPHAEL RGYNSEFGGR ASQYVAPGSV SSMFSPAAYL TELYREARNL HASDSVYHLD ERRPDLQSMT LSQQNMDTEL S TLSLSNEI LLKGIKANQS NLDSDTKVME MLSTFRPSGT IPYHDAYENV RKAIQLQDPK LEQFQKSPAV AGLMHQASLL GI NNSISPE LFNILTEEIT EANAEAIYKQ NFGDIDPACL AMPEYLKSYY NFSDEELSQF IRKYPDNELN TQKIHLLKIN KII LLSQAV NLPFLKLDEI IPEQNITPTV LGKIFLVKYY MQKYNIGTET ALILCNDSIS QYSYSNQPSQ FDRLFNTSPL NGQY FVIED TNIDLSLNST DNWHKAVLKR AFNVDDISLY RLLHIANHNN TDGKIANNIK NLSNLYMTKL LADIHQLTID ELYLL LITI GEDKINLYDI DDKELEKLIN RLDTLSNWLH TQKWSIYQLF LMTTTNYDKT LTPEIQNLLD TVYNGLQNFD KNKTKL LAA IAPYIAATLQ LPSENVAHSI LLWADKIKPS ENKITAEKFW IWLQNRDTTE LSKPPEMQEQ IIQYCHCLAQ LTMIYRS SG INENAFRLFI EKPTIFGIPD EPNKATPAHN APTLIILTRF ANWVNSLGEK ASPILTAFEN KTLTAEKLAN AMNLDANL L EQASIQAQNY KQVTKENTFS NWQSIDIILQ WTNIASNLNI SPQGISPLIA LDYIKPAQKT PTYAQWENAA IALTAGLDT QQTHTLHVFL DESRSTALSN YYIGKVANRA ASIKSRDDLY QYLLIDNQVS AEIKTTRIAE AIASIQLYVN RALENIEIHA VSDVITRQF FIDWDKYNKR YSTWAGVSQL VYYPENYIDP TMRIGQTKMM DTLLQSVSQS QLNADTVEDA FKSYLTSFEQ V ANLEVISA YHDNVNNDQG LTYFIGNSKT EVNQYYWRSV DHSKFNDGKF AANAWSEWHK IDCAINPYQS TIRPVIYKSR LY LIWLEQK ETAKQKEDNK VTTDYHYELK LAHIRYDGTW NVPITFDVDE KILALELTKS QAPGLYCAGY QGEDTLLIMF YRK KEKLDD YKTAPMQGFY IFSDMSSKDM TNEQCNSYRD NGYTHFDTNS DTNSVIRINN RYAEDYEIPS LINHSNSHDW GEYN LSQVY GGNIVINYKV TSNDLKIYIS PKLRIIHDGK EGRERIQSNL IKKYGKLGDK FIIYTSLGIN PNNSSNRFMF YPVYQ YNGN TSGLAQGRLL FHRDTSYSSK VAAWIPGAGR SLINENANIG DDCAEDSVNK PDDLKQYIYM TDSKGTATDV SGPVDI NTA ISSEKVQITI KAGKEYSLTA NKDVSVQPSP SFEEMCYQFN ALEIDGSNLN FTNNSASIDV TFTALADDGR KLGYEIF NI PVIQKVKTDN ALTLFHDENG AQYMQWGAYR IRLNTLFARQ LVERANTGID TILSMETQNI QEPMMGIGAY IELILDKY N PDIHGTNKSF KIIYGDIFKA GDHFPIYQGA LSDITQTTVK LFLPRVDNAY GNKNNLYVYA AYQKVETNFI RFVKEDNNK PATFDTTYKN GTFPGLASAR VIQTVSEPMD FSGANSLYFW ELFYYTPMMV AQRLLHEQNF DEANRWLKYV WSPSGYIVRG QIKNYHWNV RPLLENTSWN SDPLDSVDPD AVAQHDPMHY KVATFMRTLD LLMARGDHAY RQLERDTLNE AKMWYMQALH L LGNKPYLP LSSVWNDPRL DNAAATTTQK AHAYAITSLR QGTQTPALLL RSANTLTDLF LPQINDVMLS YWNKLELRLY NL RHNLSID GQPLHLPIYA TPADPKALLS AAVATSQGGG KLPESFISLW RFPHMLENAR SMVTQLIQFG STLQNIIERQ DAE SLNALL QNQAKELILT TLSIQDKTIE EIDAEKTVLE KSKAGAKSRF DNYSKLYDED VNAGERQALD MRIASQSITS GLKG LHMAA AALEMVPNIY GFAVGGTRYG AIANAIAIGG GIAAEGLLIE AEKVSQSEIW RRRRQEWEIQ RNNAEAEMKQ IDAQL KSLT VRREAAVLQK TGLKTQQEQT QAQLAFLQRK FSNQALYNWL RGRLAAIYFQ FYDLVVARCL MAEQAYRWET NDSSAR FIK PGAWQGTYAG LLAGETLMLN LAQMEDAHLK QEQRALEVER TVSLAQVYQS LGEKSFALKD KIEALLQGDK ETSAGND GN QLKLTNNTLS ATLTLQDLKL KDDYPEEMQL GKTRRIKQIS VSLPALLGPY QDVQAVLSYG GDATGLAKGC KALAVSHG L NDNGQFQLDF NDGKFLPFEG IDINDKGTFT LSFPNAASKQ KNILQMLTDI ILHIRYTILE UniProtKB: TcdA4 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average exposure time: 1.0 sec. / Average electron dose: 30.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C5 (5 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 4.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: SPHIRE / Number images used: 35857 |
| Initial angle assignment | Type: OTHER / Software - Name: SPHIRE |
| Final angle assignment | Type: OTHER / Software - Name: SPHIRE |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)