+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0065 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Transition state structure of Cpf1(Cas12a)I4 conformation | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | genome editing CRISPR ribonucleoprotein complex / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationBacillus subtilis ribonuclease / deoxyribonuclease I / deoxyribonuclease I activity / defense response to virus / lyase activity / DNA binding / RNA binding Similarity search - Function | |||||||||
| Biological species |  Francisella tularensis subsp. novicida U112 (bacteria) Francisella tularensis subsp. novicida U112 (bacteria) | |||||||||
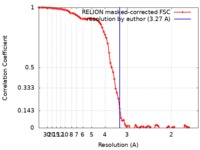
| Method | single particle reconstruction / cryo EM / Resolution: 3.27 Å | |||||||||
 Authors Authors | Mesa P / Montoya G | |||||||||
| Funding support |  Denmark, 1 items Denmark, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2018 Journal: Cell / Year: 2018Title: Conformational Activation Promotes CRISPR-Cas12a Catalysis and Resetting of the Endonuclease Activity. Authors: Stefano Stella / Pablo Mesa / Johannes Thomsen / Bijoya Paul / Pablo Alcón / Simon B Jensen / Bhargav Saligram / Matias E Moses / Nikos S Hatzakis / Guillermo Montoya /  Abstract: Cas12a, also known as Cpf1, is a type V-A CRISPR-Cas RNA-guided endonuclease that is used for genome editing based on its ability to generate specific dsDNA breaks. Here, we show cryo-EM structures ...Cas12a, also known as Cpf1, is a type V-A CRISPR-Cas RNA-guided endonuclease that is used for genome editing based on its ability to generate specific dsDNA breaks. Here, we show cryo-EM structures of intermediates of the cleavage reaction, thus visualizing three protein regions that sense the crRNA-DNA hybrid assembly triggering the catalytic activation of Cas12a. Single-molecule FRET provides the thermodynamics and kinetics of the conformational activation leading to phosphodiester bond hydrolysis. These findings illustrate why Cas12a cuts its target DNA and unleashes unspecific cleavage activity, degrading ssDNA molecules after activation. In addition, we show that other crRNAs are able to displace the R-loop inside the protein after target DNA cleavage, terminating indiscriminate ssDNA degradation. We propose a model whereby the conformational activation of the enzyme results in indiscriminate ssDNA cleavage. The displacement of the R-loop by a new crRNA molecule will reset Cas12a specificity, targeting new DNAs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0065.map.gz emd_0065.map.gz | 35.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0065-v30.xml emd-0065-v30.xml emd-0065.xml emd-0065.xml | 15.6 KB 15.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0065_fsc.xml emd_0065_fsc.xml | 8.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_0065.png emd_0065.png | 134.2 KB | ||
| Filedesc metadata |  emd-0065.cif.gz emd-0065.cif.gz | 6.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0065 http://ftp.pdbj.org/pub/emdb/structures/EMD-0065 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0065 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0065 | HTTPS FTP |
-Related structure data
| Related structure data |  6gtgMC  0061C  0062C  0063C  0064C  6gtcC  6gtdC  6gteC  6gtfC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0065.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0065.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Transition state complex I4 conformation
+Supramolecule #1: Transition state complex I4 conformation
+Supramolecule #2: CRISPR-associated endonuclease Cas12a
+Supramolecule #3: RNA (28-mer)
+Supramolecule #4: DNA
+Macromolecule #1: CRISPR-associated endonuclease Cas12a
+Macromolecule #2: RNA (40-MER)
+Macromolecule #3: DNA (32-MER)
+Macromolecule #4: DNA (5'-D(P*CP*GP*AP*GP*CP*TP*CP*GP*TP*TP*AP*GP*AP*GP*AP*AP*GP*T)-3')
+Macromolecule #5: MAGNESIUM ION
+Macromolecule #6: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)