[English] 日本語
 Yorodumi
Yorodumi- EMDB-0055: Cryo-EM reconstruction of yeast 80S ribosome in complex with mRNA... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
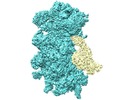
| Title | Cryo-EM reconstruction of yeast 80S ribosome in complex with mRNA, tRNA and eEF2 (GDP+AlF4/sordarin): SSU-focused map | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.3 Å | |||||||||
 Authors Authors | Pellegrino S / Yusupov M / Yusupova G / Hashem Y | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2018 Journal: J Mol Biol / Year: 2018Title: Structural Insights into the Role of Diphthamide on Elongation Factor 2 in mRNA Reading-Frame Maintenance. Authors: Simone Pellegrino / Natalia Demeshkina / Eder Mancera-Martinez / Sergey Melnikov / Angelita Simonetti / Alexander Myasnikov / Marat Yusupov / Gulnara Yusupova / Yaser Hashem /   Abstract: One of the most critical steps of protein biosynthesis is the coupled movement of mRNA, which encodes genetic information, with tRNAs on the ribosome. In eukaryotes, this process is catalyzed by a ...One of the most critical steps of protein biosynthesis is the coupled movement of mRNA, which encodes genetic information, with tRNAs on the ribosome. In eukaryotes, this process is catalyzed by a conserved G-protein, the elongation factor 2 (eEF2), which carries a unique post-translational modification, called diphthamide, found in all eukaryotic species. Here we present near-atomic resolution cryo-electron microscopy structures of yeast 80S ribosome complexes containing mRNA, tRNA and eEF2 trapped in different GTP-hydrolysis states which provide further structural insights into the role of diphthamide in the mechanism of translation fidelity in eukaryotes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|---|
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0055.map.gz emd_0055.map.gz | 12.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0055-v30.xml emd-0055-v30.xml emd-0055.xml emd-0055.xml | 22.2 KB 22.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0055.png emd_0055.png | 194.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0055 http://ftp.pdbj.org/pub/emdb/structures/EMD-0055 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0055 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0055 | HTTPS FTP |
-Related structure data
| Related structure data |  0047C  0048C  0049C  6gq1C  6gqbC  6gqvC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0055.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0055.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : 80S ribosome in complex with mRNA, tRNA and eEF2 in presence of G...
| Entire | Name: 80S ribosome in complex with mRNA, tRNA and eEF2 in presence of GDP, AlF4 and anti-fungal drug (sordarin) |
|---|---|
| Components |
|
-Supramolecule #1: 80S ribosome in complex with mRNA, tRNA and eEF2 in presence of G...
| Supramolecule | Name: 80S ribosome in complex with mRNA, tRNA and eEF2 in presence of GDP, AlF4 and anti-fungal drug (sordarin) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Molecular weight | Theoretical: 3.4 MDa |
-Supramolecule #2: 80S ribosome
| Supramolecule | Name: 80S ribosome / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#81, #83-#84 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: mRNA
| Supramolecule | Name: mRNA / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #82 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism: synthetic construct (others) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blot force 4, blot waiting time 30 s. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average exposure time: 1.5 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 4.3 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 77200 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















