+Search query
-Structure paper
| Title | Structural insights into the G-protein subtype selectivity revealed by human sphingosine-1-phosphate receptor 3-G complexes. |
|---|---|
| Journal, issue, pages | Proc Natl Acad Sci U S A, Vol. 122, Issue 47, Page e2507421122, Year 2025 |
| Publish date | Nov 25, 2025 |
 Authors Authors | Momono Yamauchi / Dohyun Im / Shintaro Maeda / Tatsuya Ikuta / Masayasu Toyomoto / Hidetsugu Asada / Yukihiko Sugita / Jun-Ichi Kishikawa / Takeshi Noda / Takayuki Kato / Asuka Inoue / So Iwata / Masatoshi Hagiwara /  |
| PubMed Abstract | Sphingosine-1-phosphate (S1P) is one of the most extensively studied bioactive lipids that transduces signals via the S1P receptor (S1PR) family (S1PR1-5), a class of G-protein-coupled receptors ...Sphingosine-1-phosphate (S1P) is one of the most extensively studied bioactive lipids that transduces signals via the S1P receptor (S1PR) family (S1PR1-5), a class of G-protein-coupled receptors (GPCRs), to regulate immune cell migration, vascular permeability, and pain modulation. However, the mechanism for achieving specificity in downstream signaling remains poorly understood. Here, we present cryogenic electron microscopic structures of the S1PR3-G complex bound to endogenous agonists: d18:1 S1P or d16:1 S1P. Both agonists shared the same binding pocket and binding mode despite the different signaling intensities of the S1PR3-G signal pathway. By comparing the structures of two agonist-bound complexes, combined with mutagenesis studies, we identified key amino acids, Phe119 and Arg136, that play crucial roles in differential agonist recognition and receptor activation. Furthermore, structural comparisons with previously determined S1PR3-G complex or G-protein-free S1PR3 structures, along with mutagenesis analysis, revealed dynamic intracellular loop 2 conformations and specific amino acid interactions that contribute to G-protein selectivity. Notably, we identified amino acids at the 34.50 and 34.53 positions within ICL2 as critical for specific interactions with G proteins. These findings provide better understanding of the mechanism of GPCR activation and unique perspectives that can be applied to other class A GPCRs, leading to the possibility of optimized drug development. |
 External links External links |  Proc Natl Acad Sci U S A / Proc Natl Acad Sci U S A /  PubMed:41252158 / PubMed:41252158 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.25 - 3.73 Å |
| Structure data | EMDB-62868, PDB-9l74: EMDB-66136, PDB-9wp9: |
| Chemicals |  PDB-1l69: 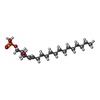 ChemComp-S1P: |
| Source |
|
 Keywords Keywords | MEMBRANE PROTEIN / GPCR / SBDD / lipid |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers







 homo sapiens (human)
homo sapiens (human)