+検索条件
-Structure paper
| タイトル | Structural and functional insights into Spns2-mediated transport of sphingosine-1-phosphate. |
|---|---|
| ジャーナル・号・ページ | Cell, Vol. 186, Issue 12, Page 2644-2655.e16, Year 2023 |
| 掲載日 | 2023年6月8日 |
 著者 著者 | Hongwen Chen / Shahbaz Ahmed / Hongtu Zhao / Nadia Elghobashi-Meinhardt / Yaxin Dai / Jae Hun Kim / Jeffrey G McDonald / Xiaochun Li / Chia-Hsueh Lee /   |
| PubMed 要旨 | Sphingosine-1-phosphate (S1P) is an important signaling sphingolipid that regulates the immune system, angiogenesis, auditory function, and epithelial and endothelial barrier integrity. Spinster ...Sphingosine-1-phosphate (S1P) is an important signaling sphingolipid that regulates the immune system, angiogenesis, auditory function, and epithelial and endothelial barrier integrity. Spinster homolog 2 (Spns2) is an S1P transporter that exports S1P to initiate lipid signaling cascades. Modulating Spns2 activity can be beneficial in treatments of cancer, inflammation, and immune diseases. However, the transport mechanism of Spns2 and its inhibition remain unclear. Here, we present six cryo-EM structures of human Spns2 in lipid nanodiscs, including two functionally relevant intermediate conformations that link the inward- and outward-facing states, to reveal the structural basis of the S1P transport cycle. Functional analyses suggest that Spns2 exports S1P via facilitated diffusion, a mechanism distinct from other MFS lipid transporters. Finally, we show that the Spns2 inhibitor 16d attenuates the transport activity by locking Spns2 in the inward-facing state. Our work sheds light on Spns2-mediated S1P transport and aids the development of advanced Spns2 inhibitors. |
 リンク リンク |  Cell / Cell /  PubMed:37224812 / PubMed:37224812 /  PubMed Central PubMed Central |
| 手法 | EM (単粒子) |
| 解像度 | 2.93 - 4.17 Å |
| 構造データ | EMDB-28650, PDB-8ex4: EMDB-28651, PDB-8ex5: EMDB-28652, PDB-8ex6: EMDB-28653, PDB-8ex7: EMDB-28654, PDB-8ex8: EMDB-29860, PDB-8g92: |
| 化合物 | 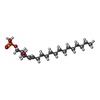 ChemComp-S1P: 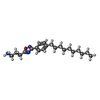 ChemComp-YUX: |
| 由来 |
|
 キーワード キーワード | MEMBRANE PROTEIN / transporter / IMMUNE SYSTEM / SPNS2 / Sphingosine-1-phosphate / major facilitator superfamily |
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア 万見文献について
万見文献について















 homo sapiens (ヒト)
homo sapiens (ヒト)