+Search query
-Structure paper
| Title | The molybdenum storage protein forms and deposits distinct polynuclear tungsten oxygen aggregates. |
|---|---|
| Journal, issue, pages | J Inorg Biochem, Vol. 234, Page 111904, Year 2022 |
| Publish date | Jun 16, 2022 |
 Authors Authors | Iram Aziz / Susann Kaltwasser / Kanwal Kayastha / Radhika Khera / Janet Vonck / Ulrich Ermler /  |
| PubMed Abstract | Some N-fixing bacteria store Mo to maintain the formation of the vital FeMo-cofactor dependent nitrogenase under Mo depleting conditions. The Mo storage protein (MoSto), developed for this purpose, ...Some N-fixing bacteria store Mo to maintain the formation of the vital FeMo-cofactor dependent nitrogenase under Mo depleting conditions. The Mo storage protein (MoSto), developed for this purpose, has the unique capability to compactly deposit molybdate as polyoxometalate (POM) clusters in a (αβ) hexameric cage; the same occurs with the physicochemically related tungstate. To explore the structural diversity of W-based POM clusters, MoSto loaded under different conditions with tungstate and two site-specifically modified MoSto variants were structurally characterized by X-ray crystallography or single-particle cryo-EM. The MoSto cage contains five major locations for POM clusters occupied among others by heptanuclear, Keggin ion and even Dawson-like species also found in bulk solvent under defined conditions. We found both lacunary derivatives of these archetypical POM clusters with missing WO units at positions exposed to bulk solvent and expanded derivatives with additional WO units next to protecting polypeptide segments or other POM clusters. The cryo-EM map, unexpectedly, reveals a POM cluster in the cage center anchored to the wall by a WO linker. Interestingly, distinct POM cluster structures can originate from identical, highly occupied core fragments of three to seven WO units that partly correspond to those found in MoSto loaded with molybdate. These core fragments are firmly bound to the complementary protein template in contrast to the more variable, less occupied residual parts of the visible POM clusters. Due to their higher stability, W-based POM clusters are, on average, larger and more diverse than their Mo-based counterparts. |
 External links External links |  J Inorg Biochem / J Inorg Biochem /  PubMed:35779405 PubMed:35779405 |
| Methods | EM (single particle) / X-ray diffraction |
| Resolution | 1.4 - 2.58 Å |
| Structure data | EMDB-14522, PDB-7z5j:  PDB-7zqq:  PDB-7zr4:  PDB-7zse: |
| Chemicals |  ChemComp-MOO:  ChemComp-ATP:  ChemComp-MG:  ChemComp-IWL:  ChemComp-IV9:  ChemComp-IWO:  ChemComp-IWZ:  ChemComp-IHW: 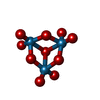 ChemComp-IX3: 
ChemComp-UNL:  ChemComp-CL:  ChemComp-PO4:  ChemComp-HOH: |
| Source |
|
 Keywords Keywords | METAL BINDING PROTEIN / Metal storage protein / polytungstate / Keggin ion / EM / polyoxotungstate / metal storage / molybdenum storage / polyoxotungstates / Molybdenum storage protein / polyoxotungstate clusters |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers





 azotobacter vinelandii dj (bacteria)
azotobacter vinelandii dj (bacteria)