+Search query
-Structure paper
| Title | Structural insights into auxin recognition and efflux by Arabidopsis PIN1. |
|---|---|
| Journal, issue, pages | Nature, Vol. 609, Issue 7927, Page 611-615, Year 2022 |
| Publish date | Aug 2, 2022 |
 Authors Authors | Zhisen Yang / Jing Xia / Jingjing Hong / Chenxi Zhang / Hong Wei / Wei Ying / Chunqiao Sun / Lianghanxiao Sun / Yanbo Mao / Yongxiang Gao / Shutang Tan / Jiří Friml / Dianfan Li / Xin Liu / Linfeng Sun /   |
| PubMed Abstract | Polar auxin transport is unique to plants and coordinates their growth and development. The PIN-FORMED (PIN) auxin transporters exhibit highly asymmetrical localizations at the plasma membrane and ...Polar auxin transport is unique to plants and coordinates their growth and development. The PIN-FORMED (PIN) auxin transporters exhibit highly asymmetrical localizations at the plasma membrane and drive polar auxin transport; however, their structures and transport mechanisms remain largely unknown. Here, we report three inward-facing conformation structures of Arabidopsis thaliana PIN1: the apo state, bound to the natural auxin indole-3-acetic acid (IAA), and in complex with the polar auxin transport inhibitor N-1-naphthylphthalamic acid (NPA). The transmembrane domain of PIN1 shares a conserved NhaA fold. In the substrate-bound structure, IAA is coordinated by both hydrophobic stacking and hydrogen bonding. NPA competes with IAA for the same site at the intracellular pocket, but with a much higher affinity. These findings inform our understanding of the substrate recognition and transport mechanisms of PINs and set up a framework for future research on directional auxin transport, one of the most crucial processes underlying plant development. |
 External links External links |  Nature / Nature /  PubMed:35917925 / PubMed:35917925 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.1 - 3.3 Å |
| Structure data | EMDB-33691, PDB-7y9t: EMDB-33692, PDB-7y9u: EMDB-33693, PDB-7y9v: |
| Chemicals |  ChemComp-E7O: 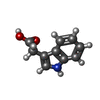 ChemComp-IAC:  ChemComp-HOH: |
| Source |
|
 Keywords Keywords | TRANSPORT PROTEIN |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers











