+Search query
-Structure paper
| Title | Broadly protective murine monoclonal antibodies against influenza B virus target highly conserved neuraminidase epitopes. |
|---|---|
| Journal, issue, pages | Nat Microbiol, Vol. 2, Issue 10, Page 1415-1424, Year 2017 |
| Publish date | Aug 21, 2017 |
 Authors Authors | Teddy John Wohlbold / Kira A Podolsky / Veronika Chromikova / Ericka Kirkpatrick / Veronica Falconieri / Philip Meade / Fatima Amanat / Jessica Tan / Benjamin R tenOever / Gene S Tan / Sriram Subramaniam / Peter Palese / Florian Krammer /  |
| PubMed Abstract | A substantial proportion of influenza-related childhood deaths are due to infection with influenza B viruses, which co-circulate in the human population as two antigenically distinct lineages defined ...A substantial proportion of influenza-related childhood deaths are due to infection with influenza B viruses, which co-circulate in the human population as two antigenically distinct lineages defined by the immunodominant receptor binding protein, haemagglutinin. While broadly cross-reactive, protective monoclonal antibodies against the haemagglutinin of influenza B viruses have been described, none targeting the neuraminidase, the second most abundant viral glycoprotein, have been reported. Here, we analyse a panel of five murine anti-neuraminidase monoclonal antibodies that demonstrate broad binding, neuraminidase inhibition, in vitro antibody-dependent cell-mediated cytotoxicity and in vivo protection against influenza B viruses belonging to both haemagglutinin lineages and spanning over 70 years of antigenic drift. Electron microscopic analysis of two neuraminidase-antibody complexes shows that the conserved neuraminidase epitopes are located on the head of the molecule and that they are distinct from the enzymatic active site. In the mouse model, one therapeutic dose of antibody 1F2 was more protective than the current standard of treatment, oseltamivir, given twice daily for six days. |
 External links External links |  Nat Microbiol / Nat Microbiol /  PubMed:28827718 / PubMed:28827718 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 25.0 Å |
| Structure data |  EMDB-8768: 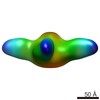 EMDB-8769:  EMDB-8770: |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers



 Influenza B virus (B/Malaysia/2506/2004)
Influenza B virus (B/Malaysia/2506/2004)