+Search query
-Structure paper
| Title | Metabolite-gated vascular contractility switch: OXGR1 activation mechanism enables agonist therapy for rosacea erythema. |
|---|---|
| Journal, issue, pages | Cell, Vol. 189, Issue 7, Page 1990-2006.e30, Year 2026 |
| Publish date | Apr 2, 2026 |
 Authors Authors | Wenqin Xiao / Yan Zhu / Xinjie Tang / Kongkai Zhu / Weifeng Zhang / Mengting Chen / Kui Cai / San Xu / Zheng Wu / Mei Wang / Jiayi Liu / Linglong Long / Zixin Tan / Aike Wu / Songqi Zhou / Zhixiang Zhao / Yan Tang / Yingxue Huang / Ben Wang / Fangfen Liu / Qian Wang / Fan Yang / Dan Jian / Wei Shi / Hongfu Xie / Xiang Chen / Lulu Guo / Zhili Deng / Jinpeng Sun / Ji Li /  |
| PubMed Abstract | Rosacea, an inflammatory skin disorder, poses a dilemma owing to limited effectiveness of treatments for pathological vasodilation-mediated erythema. Here, we identify oxoglutaric acid (α-KG) as a ...Rosacea, an inflammatory skin disorder, poses a dilemma owing to limited effectiveness of treatments for pathological vasodilation-mediated erythema. Here, we identify oxoglutaric acid (α-KG) as a rosacea-associated metabolite elevated in patients and correlated with erythema severity. Exogenous α-KG administration ameliorates rosacea-like manifestations in murine models. Mechanistically, α-KG activates OXGR1, a vascular smooth muscle cell (VSMC)-enriched G protein-coupled receptor (GPCR) to induce Gq signaling and enhance MYL9 phosphorylation, promoting VSMC contraction and limiting vasodilation. Cryo-electron microscopy (cryo-EM) structures of OXGR1-Gq complexes bound to α-KG or itaconate reveal a specific bipartite-acid pocket recognizing its endogenous agonist and an activation mechanism distinct from classical GPCRs. Building on these structures, we developed A-1, a synthetic selective OXGR1 agonist that mitigates erythema and inflammation with efficacy comparable to first-line therapy while offering enhanced safety in rosacea-like models. These findings link a metabolite to vascular dysfunction and nominate OXGR1 agonism for precision treatment of erythema and vascular disorders. |
 External links External links |  Cell / Cell /  PubMed:41791372 PubMed:41791372 |
| Methods | EM (single particle) |
| Resolution | 2.4 - 2.7 Å |
| Structure data | EMDB-65192, PDB-9vmn: EMDB-65193, PDB-9vmo: EMDB-65194, PDB-9vmp: EMDB-65222, PDB-9vo2: |
| Chemicals |  ChemComp-AKG: 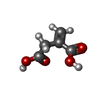 ChemComp-ITN:  PDB-1e2r: |
| Source |
|
 Keywords Keywords | MEMBRANE PROTEIN/IMMUNE SYSTEM / OXGR1 / Gq / a-KG / MEMBRANE PROTEIN / MEMBRANE PROTEIN-IMMUNE SYSTEM complex / ITA / A-1 / constitutive active |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers











 homo sapiens (human)
homo sapiens (human)