+Search query
-Structure paper
| Title | Structural insights into ligand recognition and selectivity of the human hydroxycarboxylic acid receptor HCAR2. |
|---|---|
| Journal, issue, pages | Cell Discov, Vol. 9, Issue 1, Page 118, Year 2023 |
| Publish date | Nov 28, 2023 |
 Authors Authors | Xin Pan / Fang Ye / Peiruo Ning / Zhiyi Zhang / Xinyu Li / Binghao Zhang / Qian Wang / Geng Chen / Wei Gao / Chen Qiu / Zhangsong Wu / Jiancheng Li / Lizhe Zhu / Jiang Xia / Kaizheng Gong / Yang Du /  |
| PubMed Abstract | Hydroxycarboxylic acid receptor 2 (HCAR2) belongs to the family of class A G protein-coupled receptors with key roles in regulating lipolysis and free fatty acid formation in humans. It is deeply ...Hydroxycarboxylic acid receptor 2 (HCAR2) belongs to the family of class A G protein-coupled receptors with key roles in regulating lipolysis and free fatty acid formation in humans. It is deeply involved in many pathophysiological processes and serves as an attractive target for the treatment of cardiovascular, neoplastic, autoimmune, neurodegenerative, inflammatory, and metabolic diseases. Here, we report four cryo-EM structures of human HCAR2-Gi1 complexes with or without agonists, including the drugs niacin (2.69 Å) and acipimox (3.23 Å), the highly subtype-specific agonist MK-6892 (3.25 Å), and apo form (3.28 Å). Combined with molecular dynamics simulation and functional analysis, we have revealed the recognition mechanism of HCAR2 for different agonists and summarized the general pharmacophore features of HCAR2 agonists, which are based on three key residues R111, S179, and Y284. Notably, the MK-6892-HCAR2 structure shows an extended binding pocket relative to other agonist-bound HCAR2 complexes. In addition, the key residues that determine the ligand selectivity between the HCAR2 and HCAR3 are also illuminated. Our findings provide structural insights into the ligand recognition, selectivity, activation, and G protein coupling mechanism of HCAR2, which shed light on the design of new HCAR2-targeting drugs for greater efficacy, higher selectivity, and fewer or no side effects. |
 External links External links |  Cell Discov / Cell Discov /  PubMed:38012147 / PubMed:38012147 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.69 - 3.28 Å |
| Structure data | EMDB-35463, PDB-8ij3: EMDB-35483, PDB-8ija: EMDB-35484, PDB-8ijb: EMDB-35485, PDB-8ijd: |
| Chemicals |  ChemComp-NIO: 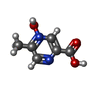 ChemComp-OJX:  ChemComp-FI7: |
| Source |
|
 Keywords Keywords | MEMBRANE PROTEIN / complex |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers











 homo sapiens (human)
homo sapiens (human)