+検索条件
-Structure paper
| タイトル | Structure of lactococcal phage p2 baseplate and its mechanism of activation. |
|---|---|
| ジャーナル・号・ページ | Proc Natl Acad Sci U S A, Vol. 107, Issue 15, Page 6852-6857, Year 2010 |
| 掲載日 | 2010年4月13日 |
 著者 著者 | Giuliano Sciara / Cecilia Bebeacua / Patrick Bron / Denise Tremblay / Miguel Ortiz-Lombardia / Julie Lichière / Marin van Heel / Valérie Campanacci / Sylvain Moineau / Christian Cambillau /  |
| PubMed 要旨 | Siphoviridae is the most abundant viral family on earth which infects bacteria as well as archaea. All known siphophages infecting gram+ Lactococcus lactis possess a baseplate at the tip of their ...Siphoviridae is the most abundant viral family on earth which infects bacteria as well as archaea. All known siphophages infecting gram+ Lactococcus lactis possess a baseplate at the tip of their tail involved in host recognition and attachment. Here, we report analysis of the p2 phage baseplate structure by X-ray crystallography and electron microscopy and propose a mechanism for the baseplate activation during attachment to the host cell. This approximately 1 MDa, Escherichia coli-expressed baseplate is composed of three protein species, including six trimers of the receptor-binding protein (RBP). RBPs host-recognition domains point upwards, towards the capsid, in agreement with the electron-microscopy map of the free virion. In the presence of Ca(2+), a cation mandatory for infection, the RBPs rotated 200 degrees downwards, presenting their binding sites to the host, and a channel opens at the bottom of the baseplate for DNA passage. These conformational changes reveal a novel siphophage activation and host-recognition mechanism leading ultimately to DNA ejection. |
 リンク リンク |  Proc Natl Acad Sci U S A / Proc Natl Acad Sci U S A /  PubMed:20351260 / PubMed:20351260 /  PubMed Central PubMed Central |
| 手法 | EM (単粒子) / X線回折 |
| 解像度 | 2.6 - 26.0 Å |
| 構造データ |  EMDB-1699: 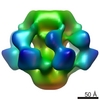 EMDB-1706:  PDB-2wzp:  PDB-2x53:  PDB-4v5i: |
| 化合物 |  ChemComp-HOH:  ChemComp-SR:  ChemComp-CA: |
| 由来 |
|
 キーワード キーワード | VIRAL PROTEIN / BASEPLATE / SIPHOVIRIDAE / LACTOCOCCUS LACTIS |
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア 万見文献について
万見文献について



 lactococcus phage p2 (ウイルス)
lactococcus phage p2 (ウイルス)
