+Search query
-Structure paper
| Title | Asymmetric Structure of Podophage GP4 Reveals a Novel Architecture of Three Types of Tail Fibers. |
|---|---|
| Journal, issue, pages | J Mol Biol, Vol. 435, Issue 20, Page 168258, Year 2023 |
| Publish date | Oct 15, 2023 |
 Authors Authors | Jing Zheng / Wenyuan Chen / Hao Xiao / Fan Yang / Jingdong Song / Lingpeng Cheng / Hongrong Liu /  |
| PubMed Abstract | Bacteriophage tail fibers (or called tail spikes) play a critical role in the early stage of infection by binding to the bacterial surface. Podophages with known structures usually possess one or two ...Bacteriophage tail fibers (or called tail spikes) play a critical role in the early stage of infection by binding to the bacterial surface. Podophages with known structures usually possess one or two types of fibers. Here, we resolved an asymmetric structure of the podophage GP4 to near-atomic resolution by cryo-EM. Our structure revealed a symmetry-mismatch relationship between the components of the GP4 tail with previously unseen topologies. In detail, two dodecameric adaptors (adaptors I and II), a hexameric nozzle, and a tail needle form a conserved tail body connected to a dodecameric portal occupying a unique vertex of the icosahedral head. However, five chain-like extended fibers (fiber I) and five tulip-like short fibers (fiber II) are anchored to a 15-fold symmetric fiber-tail adaptor, encircling the adaptor I, and six bamboo-like trimeric fibers (fiber III) are connected to the nozzle. Five fibers I, each composed of five dimers of the protein gp80 linked by an elongated rope protein, are attached to the five edges of the tail vertex of the icosahedral head. In this study, we identified a new structure of the podophage with three types of tail fibers, and such phages with different types of fibers may have a broad host range and/or infect host cells with considerably high efficiency, providing evolutionary advantages in harsh environments. |
 External links External links |  J Mol Biol / J Mol Biol /  PubMed:37660940 PubMed:37660940 |
| Methods | EM (single particle) |
| Resolution | 3.8 - 7.3 Å |
| Structure data | EMDB-36462, PDB-8jou: EMDB-36463, PDB-8jov: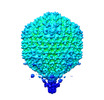  EMDB-36464: Asymmetric reconstruction map of phage GP4 |
| Source |
|
 Keywords Keywords | VIRAL PROTEIN / Complex |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers



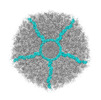



 ralstonia phage gp4 (virus)
ralstonia phage gp4 (virus)