+Search query
-Structure paper
| Title | Molecular architecture of mammalian nitric oxide synthases. |
|---|---|
| Journal, issue, pages | Proc Natl Acad Sci U S A, Vol. 111, Issue 35, Page E3614-E3623, Year 2014 |
| Publish date | Sep 2, 2014 |
 Authors Authors | Melody G Campbell / Brian C Smith / Clinton S Potter / Bridget Carragher / Michael A Marletta /  |
| PubMed Abstract | NOSs are homodimeric multidomain enzymes responsible for producing NO. In mammals, NO acts as an intercellular messenger in a variety of signaling reactions, as well as a cytotoxin in the innate ...NOSs are homodimeric multidomain enzymes responsible for producing NO. In mammals, NO acts as an intercellular messenger in a variety of signaling reactions, as well as a cytotoxin in the innate immune response. Mammals possess three NOS isoforms--inducible, endothelial, and neuronal NOS--that are composed of an N-terminal oxidase domain and a C-terminal reductase domain. Calmodulin (CaM) activates NO synthesis by binding to the helical region connecting these two domains. Although crystal structures of isolated domains have been reported, no structure is available for full-length NOS. We used high-throughput single-particle EM to obtain the structures and higher-order domain organization of all three NOS holoenzymes. The structures of inducible, endothelial, and neuronal NOS with and without CaM bound are similar, consisting of a dimerized oxidase domain flanked by two separated reductase domains. NOS isoforms adopt many conformations enabled by three flexible linkers. These conformations represent snapshots of the continuous electron transfer pathway from the reductase domain to the oxidase domain, which reveal that only a single reductase domain participates in electron transfer at a time, and that CaM activates NOS by constraining rotational motions and by directly binding to the oxidase domain. Direct visualization of these large conformational changes induced during electron transfer provides significant insight into the molecular underpinnings governing NO formation. |
 External links External links |  Proc Natl Acad Sci U S A / Proc Natl Acad Sci U S A /  PubMed:25125509 / PubMed:25125509 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 60.0 - 77.0 Å |
| Structure data |  EMDB-2718:  EMDB-2719:  EMDB-2720:  EMDB-2721:  EMDB-2722:  EMDB-2723:  EMDB-2724:  EMDB-2725:  EMDB-2726:  EMDB-2727:  EMDB-2728:  EMDB-2729:  EMDB-2730: 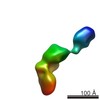 EMDB-2731:  EMDB-2732:  EMDB-2733:  EMDB-2734:  EMDB-2735:  EMDB-2736:  EMDB-2737:  EMDB-2738:  EMDB-2739:  EMDB-2740:  EMDB-2741:  EMDB-2742:  EMDB-2743:  EMDB-2744:  EMDB-2745:  EMDB-2746:  EMDB-2747:  EMDB-2748:  EMDB-2749: |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers




