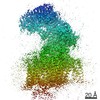+Search query
-Structure paper
| Title | Molecular Basis for Ligand Modulation of a Mammalian Voltage-Gated Ca Channel. |
|---|---|
| Journal, issue, pages | Cell, Vol. 177, Issue 6, Page 1495-1506.e12, Year 2019 |
| Publish date | May 30, 2019 |
 Authors Authors | Yanyu Zhao / Gaoxingyu Huang / Jianping Wu / Qiurong Wu / Shuai Gao / Zhen Yan / Jianlin Lei / Nieng Yan /   |
| PubMed Abstract | The L-type voltage-gated Ca (Ca) channels are modulated by various compounds exemplified by 1,4-dihydropyridines (DHP), benzothiazepines (BTZ), and phenylalkylamines (PAA), many of which have been ...The L-type voltage-gated Ca (Ca) channels are modulated by various compounds exemplified by 1,4-dihydropyridines (DHP), benzothiazepines (BTZ), and phenylalkylamines (PAA), many of which have been used for characterizing channel properties and for treatment of hypertension and other disorders. Here, we report the cryoelectron microscopy (cryo-EM) structures of Ca1.1 in complex with archetypal antagonistic drugs, nifedipine, diltiazem, and verapamil, at resolutions of 2.9 Å, 3.0 Å, and 2.7 Å, respectively, and with a DHP agonist Bay K 8644 at 2.8 Å. Diltiazem and verapamil traverse the central cavity of the pore domain, directly blocking ion permeation. Although nifedipine and Bay K 8644 occupy the same fenestration site at the interface of repeats III and IV, the coordination details support previous functional observations that Bay K 8644 is less favored in the inactivated state. These structures elucidate the modes of action of different Ca ligands and establish a framework for structure-guided drug discovery. |
 External links External links |  Cell / Cell /  PubMed:31150622 PubMed:31150622 |
| Methods | EM (single particle) |
| Resolution | 2.6 - 2.9 Å |
| Structure data | EMDB-9866, PDB-6jp5: |
| Chemicals |  ChemComp-NAG:  ChemComp-CA:  ChemComp-3PE: 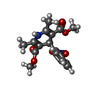 ChemComp-C5U:  ChemComp-PC1: 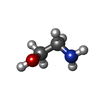 ChemComp-ETA: 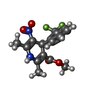 ChemComp-C8U:  ChemComp-9Z9:  ChemComp-4YH:  ChemComp-C9F: |
| Source |
|
 Keywords Keywords | MEMBRANE PROTEIN / Membrane protein complex modulated by FDA approved drug. |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers