+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8630 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
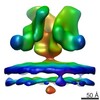
| Title | Ebola GP spike in situ in viral membrane | |||||||||
 Map data Map data | Ebola GP spike in situ in viral envelope | |||||||||
 Sample Sample | Ebola GP spike != Ebola virus Ebola GP spike
| |||||||||
| Function / homology | Filoviruses glycoprotein, extracellular domain / Filoviruses glycoprotein / Filovirus glycoprotein / membrane => GO:0016020 / extracellular region / GP Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 11.0 Å | |||||||||
 Authors Authors | Booth TF / Beniac DR | |||||||||
 Citation Citation |  Journal: Sci Rep / Year: 2017 Journal: Sci Rep / Year: 2017Title: Structure of the Ebola virus glycoprotein spike within the virion envelope at 11 Å resolution. Authors: Daniel R Beniac / Timothy F Booth /  Abstract: We present the structure of the surface Ebola virus (EBOV) trimeric glycoprotein (GP) spike at 11 Å resolution, in situ within the viral plasma membrane of purified virus particles. GP functions ...We present the structure of the surface Ebola virus (EBOV) trimeric glycoprotein (GP) spike at 11 Å resolution, in situ within the viral plasma membrane of purified virus particles. GP functions in cellular attachment, endosomal entry, and membrane fusion to initiate infection, and is a key therapeutic target. Nevertheless, only about half of the GP molecule has yet been solved to atomic resolution, excluding the mucin-like and transmembrane domains, and some of the glycans. Fitting of the atomic resolution X-ray data from expressed, truncated deletion constructs within our 11 Å structure of the entire molecule demonstrates the relationship between the GP1-GP2 domains, the mucin-like and transmembrane domains, and the bilaminar lipid envelope. We show that the mucin-like domain covers the glycan cap and partially occludes the receptor binding sites prior to proteolytic cleavage. Our structure is also consistent with key antibody neutralisation sites on GP being accessible prior to proteolysis. Based on the findings of us and others, GP-mediated binding may create an angle of 18 degrees between the planes of viral and endosomal membranes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8630.map.gz emd_8630.map.gz | 12.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8630-v30.xml emd-8630-v30.xml emd-8630.xml emd-8630.xml | 6.8 KB 6.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8630.png emd_8630.png | 164.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8630 http://ftp.pdbj.org/pub/emdb/structures/EMD-8630 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8630 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8630 | HTTPS FTP |
-Validation report
| Summary document |  emd_8630_validation.pdf.gz emd_8630_validation.pdf.gz | 373.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_8630_full_validation.pdf.gz emd_8630_full_validation.pdf.gz | 373.5 KB | Display | |
| Data in XML |  emd_8630_validation.xml.gz emd_8630_validation.xml.gz | 5.5 KB | Display | |
| Data in CIF |  emd_8630_validation.cif.gz emd_8630_validation.cif.gz | 6.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8630 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8630 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8630 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8630 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8630.map.gz / Format: CCP4 / Size: 12.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8630.map.gz / Format: CCP4 / Size: 12.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Ebola GP spike in situ in viral envelope | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.15 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Ebola GP spike
| Entire | Name: Ebola GP spike |
|---|---|
| Components |
|
-Supramolecule #1: Ebola virus
| Supramolecule | Name: Ebola virus / type: virus / ID: 1 / Parent: 0 / NCBI-ID: 1570291 / Sci species name: Ebola virus / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 20 |
|---|---|
| Image recording | Film or detector model: FEI EAGLE (4k x 4k) / Average electron dose: 10.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 11.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 32960 |
|---|---|
| Initial angle assignment | Type: OTHER |
| Final angle assignment | Type: OTHER |
 Movie
Movie Controller
Controller



 UCSF Chimera
UCSF Chimera