[English] 日本語
 Yorodumi
Yorodumi- EMDB-1728: 70S Ribosome and Human U4U6U5 tri-snRNP Determined by Cryo-Random... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1728 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 70S Ribosome and Human U4U6U5 tri-snRNP Determined by Cryo-Random Conical Tilt | |||||||||
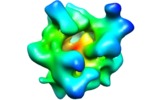
 Map data Map data | This 3D map is a weighted average cryo RCT structure of the 70S ribosome. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ribosome wRCT cryo-RCT | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 31.0 Å | |||||||||
 Authors Authors | Sander B / Golas MM / Luhrmann R / Stark H | |||||||||
 Citation Citation |  Journal: Structure / Year: 2010 Journal: Structure / Year: 2010Title: An approach for de novo structure determination of dynamic molecular assemblies by electron cryomicroscopy. Authors: Bjoern Sander / Monika M Golas / Reinhard Lührmann / Holger Stark /  Abstract: Single-particle electron cryomicroscopy is a powerful method for three-dimensional (3D) structure determination of macromolecular assemblies. Here we address the challenge of determining a 3D ...Single-particle electron cryomicroscopy is a powerful method for three-dimensional (3D) structure determination of macromolecular assemblies. Here we address the challenge of determining a 3D structure in the absence of reference models. The 3D structures are determined by alignment and weighted averaging of densities obtained by native cryo random conical tilt (RCT) reconstructions including consideration of missing data. Our weighted averaging scheme (wRCT) offers advantages for potentially heterogeneous 3D densities of low signal-to-noise ratios. Sets of aligned RCT structures can also be analyzed by multivariate statistical analysis (MSA) to provide insights into snapshots of the assemblies. The approach is used to compute 3D structures of the Escherichia coli 70S ribosome and the human U4/U6.U5 tri-snRNP under vitrified unstained cryo conditions, and to visualize by 3D MSA the L7/L12 stalk of the 70S ribosome and states of tri-snRNP. The approach thus combines de novo 3D structure determination with an analysis of compositional and conformational heterogeneity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1728.map.gz emd_1728.map.gz | 2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1728-v30.xml emd-1728-v30.xml emd-1728.xml emd-1728.xml | 6.7 KB 6.7 KB | Display Display |  EMDB header EMDB header |
| Images |  1728.gif 1728.gif 1728.tif 1728.tif | 68.7 KB 291.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1728 http://ftp.pdbj.org/pub/emdb/structures/EMD-1728 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1728 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1728 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1728.map.gz / Format: CCP4 / Size: 3.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1728.map.gz / Format: CCP4 / Size: 3.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This 3D map is a weighted average cryo RCT structure of the 70S ribosome. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.9 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : E. coli 70S ribosome
| Entire | Name: E. coli 70S ribosome |
|---|---|
| Components |
|
-Supramolecule #1000: E. coli 70S ribosome
| Supramolecule | Name: E. coli 70S ribosome / type: sample / ID: 1000 / Details: The sample was monodisperse / Number unique components: 1 |
|---|
-Supramolecule #1: Ribosome
| Supramolecule | Name: Ribosome / type: complex / ID: 1 / Name.synonym: E. coli 70S / Recombinant expression: No / Ribosome-details: ribosome-prokaryote: ALL |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM200FEG |
|---|---|
| Electron beam | Acceleration voltage: 160 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 31.0 Å / Resolution method: OTHER |
|---|
-Atomic model buiding 1
| Initial model | PDB ID:  2aw7 |
|---|---|
| Details | Protocol: Rigid Body |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
-Atomic model buiding 2
| Initial model | PDB ID:  2awb |
|---|---|
| Details | Protocol: Rigid Body |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















