[English] 日本語
 Yorodumi
Yorodumi- EMDB-8408: Cryo-EM reconstruction of the yeast kinesin-5, Cin8, bound to GDP... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8408 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM reconstruction of the yeast kinesin-5, Cin8, bound to GDP-taxol microtubules in ADP-AlFx state | |||||||||
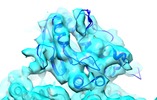
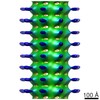
 Map data Map data | 3D helical reconstruction of Cin8 motor domain in ADP-AlFx state bound to microtubule using cryo-EM | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
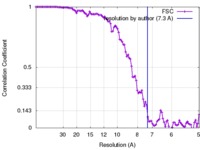
| Method | helical reconstruction / cryo EM / Resolution: 7.3 Å | |||||||||
 Authors Authors | Cha HK | |||||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2017 Journal: J Biol Chem / Year: 2017Title: The yeast kinesin-5 Cin8 interacts with the microtubule in a noncanonical manner. Authors: Kayla M Bell / Hyo Keun Cha / Charles V Sindelar / Jared C Cochran /  Abstract: Kinesin motors play central roles in establishing and maintaining the mitotic spindle during cell division. Unlike most other kinesins, Cin8, a kinesin-5 motor in can move bidirectionally along ...Kinesin motors play central roles in establishing and maintaining the mitotic spindle during cell division. Unlike most other kinesins, Cin8, a kinesin-5 motor in can move bidirectionally along microtubules, switching directionality according to biochemical conditions, a behavior that remains largely unexplained. To this end, we used biochemical rate and equilibrium constant measurements as well as cryo-electron microscopy methodologies to investigate the microtubule interactions of the Cin8 motor domain. These experiments unexpectedly revealed that, whereas Cin8 ATPase kinetics fell within measured ranges for kinesins (especially kinesin-5 proteins), approximately four motors can bind each αβ-tubulin dimer within the microtubule lattice. This result contrasted with those observations on other known kinesins, which can bind only a single "canonical" site per tubulin dimer. Competition assays with human kinesin-5 (Eg5) only partially abrogated this behavior, indicating that Cin8 binds microtubules not only at the canonical site, but also one or more separate ("noncanonical") sites. Moreover, we found that deleting the large, class-specific insert in the microtubule-binding loop 8 reverts Cin8 to one motor per αβ-tubulin in the microtubule. The novel microtubule-binding mode of Cin8 identified here provides a potential explanation for Cin8 clustering along microtubules and potentially may contribute to the mechanism for direction reversal. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8408.map.gz emd_8408.map.gz | 9.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8408-v30.xml emd-8408-v30.xml emd-8408.xml emd-8408.xml | 9.5 KB 9.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8408_fsc.xml emd_8408_fsc.xml | 11.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_8408.png emd_8408.png | 210.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8408 http://ftp.pdbj.org/pub/emdb/structures/EMD-8408 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8408 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8408 | HTTPS FTP |
-Validation report
| Summary document |  emd_8408_validation.pdf.gz emd_8408_validation.pdf.gz | 78.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_8408_full_validation.pdf.gz emd_8408_full_validation.pdf.gz | 78 KB | Display | |
| Data in XML |  emd_8408_validation.xml.gz emd_8408_validation.xml.gz | 493 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8408 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8408 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8408 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8408 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8408.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8408.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D helical reconstruction of Cin8 motor domain in ADP-AlFx state bound to microtubule using cryo-EM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.49402 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Microtubule-bound kinesin-5 Cin8 in ADP-AlFx state
| Entire | Name: Microtubule-bound kinesin-5 Cin8 in ADP-AlFx state |
|---|---|
| Components |
|
-Supramolecule #1: Microtubule-bound kinesin-5 Cin8 in ADP-AlFx state
| Supramolecule | Name: Microtubule-bound kinesin-5 Cin8 in ADP-AlFx state / type: complex / ID: 1 / Parent: 0 Details: Cin8-his monomer (residues 70-535) Taxol-stabilized microtubules |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 Component:
Details: adjusted to pH 6.8 with KOH | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.71 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller