+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5945 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Pyruvate Carboxylase tetramer in asymmetric architecture | |||||||||
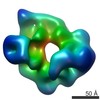
 Map data Map data | CryoEM of tetrameric Pyruvate Carboxylase in asymmetric architecture. Calculated after sorting of a catalytically active population. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | enzyme / tetrameric / biotin carboxylase | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 11.5 Å | |||||||||
 Authors Authors | Lasso G / Yu LPC / Gil D / Lazaro M / Tong L / Valle M | |||||||||
 Citation Citation |  Journal: Structure / Year: 2014 Journal: Structure / Year: 2014Title: Functional conformations for pyruvate carboxylase during catalysis explored by cryoelectron microscopy. Authors: Gorka Lasso / Linda P C Yu / David Gil / Melisa Lázaro / Liang Tong / Mikel Valle /   Abstract: The tetrameric enzyme pyruvate carboxylase (PC), a biotin-dependent carboxylase, produces oxaloacetate by two consecutive reactions that take place in distant active sites. Previous crystal ...The tetrameric enzyme pyruvate carboxylase (PC), a biotin-dependent carboxylase, produces oxaloacetate by two consecutive reactions that take place in distant active sites. Previous crystal structures revealed two different configurations for PC tetramers, the so-called symmetric and asymmetric, which were understood as characteristic molecular architectures for PC from different organisms. We have analyzed PC samples from Staphylococcus aureus while the enzyme generates oxaloacetate, expecting PC tetramers to display the conformational landscape relevant for its functioning. Using cryoelectron microscopy (cryo-EM) and sorting techniques, we detect previously defined symmetric and asymmetric architectures, demonstrating that PC maps both arrangements by large conformational changes. Furthermore, we observe that each configuration is coupled to one of the two consecutive enzymatic reactions. The findings describe the structural transitions relevant for the allosteric control of the multifunctional PC and demonstrate that by cryo-EM and classification, we can characterize freely working macromolecules. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5945.map.gz emd_5945.map.gz | 2.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5945-v30.xml emd-5945-v30.xml emd-5945.xml emd-5945.xml | 9.9 KB 9.9 KB | Display Display |  EMDB header EMDB header |
| Images |  400_5945.gif 400_5945.gif 80_5945.gif 80_5945.gif | 49.6 KB 3.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5945 http://ftp.pdbj.org/pub/emdb/structures/EMD-5945 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5945 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5945 | HTTPS FTP |
-Validation report
| Summary document |  emd_5945_validation.pdf.gz emd_5945_validation.pdf.gz | 78.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5945_full_validation.pdf.gz emd_5945_full_validation.pdf.gz | 77.7 KB | Display | |
| Data in XML |  emd_5945_validation.xml.gz emd_5945_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5945 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5945 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5945 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5945 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5945.map.gz / Format: CCP4 / Size: 25.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5945.map.gz / Format: CCP4 / Size: 25.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM of tetrameric Pyruvate Carboxylase in asymmetric architecture. Calculated after sorting of a catalytically active population. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.75 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Pyruvate Carboxylase
| Entire | Name: Pyruvate Carboxylase |
|---|---|
| Components |
|
-Supramolecule #1000: Pyruvate Carboxylase
| Supramolecule | Name: Pyruvate Carboxylase / type: sample / ID: 1000 / Oligomeric state: Homotetramer / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 520 KDa |
-Macromolecule #1: pyruvate carboxylate
| Macromolecule | Name: pyruvate carboxylate / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Oligomeric state: tetramer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 520 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.05 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 16.6 mM Tris-HCl, 1.7 mM acetyl-CoA, 15.7 mM pyruvate, 4.1 mM MgCl2, 1.7 mM DTT, 166.6 mM NaCl, 8.3 mM KHCO3, 1.66 mM ATP. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Instrument: FEI VITROBOT MARK II |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2200FSC |
|---|---|
| Specialist optics | Energy filter - Name: Omega in-column |
| Date | Jan 1, 2011 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7 µm / Number real images: 50 / Average electron dose: 15 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal magnification: 40000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| CTF correction | Details: Volume correction in defocus groups |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 11.5 Å / Resolution method: OTHER / Software - Name: Relion, Spider / Number images used: 14032 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name: MDFF |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
 Movie
Movie Controller
Controller