[English] 日本語
 Yorodumi
Yorodumi- EMDB-3508: Structural basis for ArfA-RF2 mediated translation termination on... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3508 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structural basis for ArfA-RF2 mediated translation termination on stop-codon lacking mRNAs | |||||||||
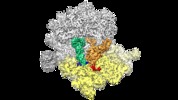
 Map data Map data | E.coli Release Factor RF2 and E.coli Alternative Ribosome-Rescue Factor ArfA bound to the 70S E.coli ribosome. | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationtranslation release factor activity, codon specific / negative regulation of cytoplasmic translational initiation / ribosomal large subunit binding / ornithine decarboxylase inhibitor activity / transcription antitermination factor activity, RNA binding / misfolded RNA binding / Group I intron splicing / RNA folding / transcriptional attenuation / endoribonuclease inhibitor activity ...translation release factor activity, codon specific / negative regulation of cytoplasmic translational initiation / ribosomal large subunit binding / ornithine decarboxylase inhibitor activity / transcription antitermination factor activity, RNA binding / misfolded RNA binding / Group I intron splicing / RNA folding / transcriptional attenuation / endoribonuclease inhibitor activity / RNA-binding transcription regulator activity / positive regulation of ribosome biogenesis / negative regulation of cytoplasmic translation / four-way junction DNA binding / translational termination / DnaA-L2 complex / translation repressor activity / negative regulation of translational initiation / negative regulation of DNA-templated DNA replication initiation / regulation of mRNA stability / rescue of stalled ribosome / mRNA regulatory element binding translation repressor activity / ribosome assembly / positive regulation of RNA splicing / assembly of large subunit precursor of preribosome / transcription elongation factor complex / cytosolic ribosome assembly / regulation of DNA-templated transcription elongation / DNA endonuclease activity / ribosomal large subunit assembly / transcription antitermination / response to reactive oxygen species / translational initiation / regulation of cell growth / DNA-templated transcription termination / maintenance of translational fidelity / response to radiation / mRNA 5'-UTR binding / large ribosomal subunit / ribosome biogenesis / ribosome binding / regulation of translation / ribosomal small subunit biogenesis / ribosomal small subunit assembly / small ribosomal subunit / small ribosomal subunit rRNA binding / transferase activity / 5S rRNA binding / large ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / molecular adaptor activity / rRNA binding / negative regulation of translation / ribosome / structural constituent of ribosome / translation / response to antibiotic / negative regulation of DNA-templated transcription / mRNA binding / DNA binding / RNA binding / zinc ion binding / membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Huter P / Mueller C / Beckert B / Arenz S / Berninghausen O / Beckmann R / Wilson ND | |||||||||
 Citation Citation |  Journal: Nature / Year: 2017 Journal: Nature / Year: 2017Title: Structural basis for ArfA-RF2-mediated translation termination on mRNAs lacking stop codons. Authors: Paul Huter / Claudia Müller / Bertrand Beckert / Stefan Arenz / Otto Berninghausen / Roland Beckmann / Daniel N Wilson /  Abstract: In bacteria, ribosomes stalled on truncated mRNAs that lack a stop codon are rescued by the transfer-messenger RNA (tmRNA), alternative rescue factor A (ArfA) or ArfB systems. Although tmRNA-ribosome ...In bacteria, ribosomes stalled on truncated mRNAs that lack a stop codon are rescued by the transfer-messenger RNA (tmRNA), alternative rescue factor A (ArfA) or ArfB systems. Although tmRNA-ribosome and ArfB-ribosome structures have been determined, how ArfA recognizes the presence of truncated mRNAs and recruits the canonical termination release factor RF2 to rescue the stalled ribosomes is unclear. Here we present a cryo-electron microscopy reconstruction of the Escherichia coli 70S ribosome stalled on a truncated mRNA in the presence of ArfA and RF2. The structure shows that the C terminus of ArfA binds within the mRNA entry channel on the small ribosomal subunit, and explains how ArfA distinguishes between ribosomes that bear truncated or full-length mRNAs. The N terminus of ArfA establishes several interactions with the decoding domain of RF2, and this finding illustrates how ArfA recruits RF2 to the stalled ribosome. Furthermore, ArfA is shown to stabilize a unique conformation of the switch loop of RF2, which mimics the canonical translation termination state by directing the catalytically important GGQ motif within domain 3 of RF2 towards the peptidyl-transferase centre of the ribosome. Thus, our structure reveals not only how ArfA recruits RF2 to the ribosome but also how it promotes an active conformation of RF2 to enable translation termination in the absence of a stop codon. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3508.map.gz emd_3508.map.gz | 17.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3508-v30.xml emd-3508-v30.xml emd-3508.xml emd-3508.xml | 69.9 KB 69.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_3508.png emd_3508.png | 249.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3508 http://ftp.pdbj.org/pub/emdb/structures/EMD-3508 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3508 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3508 | HTTPS FTP |
-Validation report
| Summary document |  emd_3508_validation.pdf.gz emd_3508_validation.pdf.gz | 281.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_3508_full_validation.pdf.gz emd_3508_full_validation.pdf.gz | 280.6 KB | Display | |
| Data in XML |  emd_3508_validation.xml.gz emd_3508_validation.xml.gz | 4.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3508 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3508 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3508 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3508 | HTTPS FTP |
-Related structure data
| Related structure data |  5mgpMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3508.map.gz / Format: CCP4 / Size: 57.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3508.map.gz / Format: CCP4 / Size: 57.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | E.coli Release Factor RF2 and E.coli Alternative Ribosome-Rescue Factor ArfA bound to the 70S E.coli ribosome. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.084 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Structural basis for ArfA-RF2 mediated translation termination 1 ...
+Supramolecule #1: Structural basis for ArfA-RF2 mediated translation termination 1 ...
+Supramolecule #2: ribosome
+Supramolecule #3: mRNA, tRNA, Peptide chain release factor 2
+Macromolecule #1: 23S ribosomal RNA
+Macromolecule #2: 5S ribosomal RNA
+Macromolecule #32: 16S ribosomal RNA
+Macromolecule #53: mRNA
+Macromolecule #55: P-site tRNA
+Macromolecule #3: 50S ribosomal protein L2
+Macromolecule #4: 50S ribosomal protein L3
+Macromolecule #5: 50S ribosomal protein L4
+Macromolecule #6: 50S ribosomal protein L5
+Macromolecule #7: 50S ribosomal protein L6
+Macromolecule #8: 50S ribosomal protein L9
+Macromolecule #9: 50S ribosomal protein L13
+Macromolecule #10: 50S ribosomal protein L14
+Macromolecule #11: 50S ribosomal protein L15
+Macromolecule #12: 50S ribosomal protein L16
+Macromolecule #13: 50S ribosomal protein L17
+Macromolecule #14: 50S ribosomal protein L18
+Macromolecule #15: 50S ribosomal protein L19
+Macromolecule #16: 50S ribosomal protein L20
+Macromolecule #17: 50S ribosomal protein L21
+Macromolecule #18: 50S ribosomal protein L22
+Macromolecule #19: 50S ribosomal protein L23
+Macromolecule #20: 50S ribosomal protein L24
+Macromolecule #21: 50S ribosomal protein L25
+Macromolecule #22: 50S ribosomal protein L27
+Macromolecule #23: 50S ribosomal protein L28
+Macromolecule #24: 50S ribosomal protein L29
+Macromolecule #25: 50S ribosomal protein L30
+Macromolecule #26: 50S ribosomal protein L32
+Macromolecule #27: 50S ribosomal protein L33
+Macromolecule #28: 50S ribosomal protein L34
+Macromolecule #29: 50S ribosomal protein L35
+Macromolecule #30: 50S ribosomal protein L36
+Macromolecule #31: 50S ribosomal protein L31
+Macromolecule #33: 30S ribosomal protein S2
+Macromolecule #34: 30S ribosomal protein S3
+Macromolecule #35: 30S ribosomal protein S4
+Macromolecule #36: 30S ribosomal protein S5
+Macromolecule #37: 30S ribosomal protein S6
+Macromolecule #38: 30S ribosomal protein S7
+Macromolecule #39: 30S ribosomal protein S8
+Macromolecule #40: 30S ribosomal protein S9
+Macromolecule #41: 30S ribosomal protein S10
+Macromolecule #42: 30S ribosomal protein S11
+Macromolecule #43: 30S ribosomal protein S12
+Macromolecule #44: 30S ribosomal protein S13
+Macromolecule #45: 30S ribosomal protein S14
+Macromolecule #46: 30S ribosomal protein S15
+Macromolecule #47: 30S ribosomal protein S16
+Macromolecule #48: 30S ribosomal protein S17
+Macromolecule #49: 30S ribosomal protein S18
+Macromolecule #50: 30S ribosomal protein S19
+Macromolecule #51: 30S ribosomal protein S20
+Macromolecule #52: 30S ribosomal protein S21
+Macromolecule #54: Alternative ribosome-rescue factor A
+Macromolecule #56: Peptide chain release factor 2
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average electron dose: 2.5 e/Å2 |
| Electron beam | Acceleration voltage: 302 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 3.1 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: FREALIGN (ver. 9.11) / Number images used: 69089 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller