[English] 日本語
 Yorodumi
Yorodumi- EMDB-9884: Structure of human soluble guanylate cyclase in the heme oxidised... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9884 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of human soluble guanylate cyclase in the heme oxidised state | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | soluble guanylate cyclase / SIGNALING PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationretrograde trans-synaptic signaling by nitric oxide, modulating synaptic transmission / cytidylate cyclase activity / guanylate cyclase complex, soluble / guanylate cyclase / cGMP biosynthetic process / guanylate cyclase activity / presynaptic active zone cytoplasmic component / response to oxygen levels / nitric oxide binding / relaxation of vascular associated smooth muscle ...retrograde trans-synaptic signaling by nitric oxide, modulating synaptic transmission / cytidylate cyclase activity / guanylate cyclase complex, soluble / guanylate cyclase / cGMP biosynthetic process / guanylate cyclase activity / presynaptic active zone cytoplasmic component / response to oxygen levels / nitric oxide binding / relaxation of vascular associated smooth muscle / Nitric oxide stimulates guanylate cyclase / adenylate cyclase activity / : / blood circulation / : / Smooth Muscle Contraction / : / nitric oxide-cGMP-mediated signaling / cellular response to nitric oxide / Hsp90 protein binding / GABA-ergic synapse / regulation of blood pressure / signaling receptor activity / heme binding / GTP binding / protein-containing complex binding / glutamatergic synapse / metal ion binding / cytosol Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||||||||
 Authors Authors | Chen L / Kang Y | |||||||||||||||
| Funding support |  China, 4 items China, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2019 Journal: Nature / Year: 2019Title: Structural insights into the mechanism of human soluble guanylate cyclase. Authors: Yunlu Kang / Rui Liu / Jing-Xiang Wu / Lei Chen /  Abstract: Soluble guanylate cyclase (sGC) is the primary sensor of nitric oxide. It has a central role in nitric oxide signalling and has been implicated in many essential physiological processes and disease ...Soluble guanylate cyclase (sGC) is the primary sensor of nitric oxide. It has a central role in nitric oxide signalling and has been implicated in many essential physiological processes and disease conditions. The binding of nitric oxide boosts the enzymatic activity of sGC. However, the mechanism by which nitric oxide activates the enzyme is unclear. Here we report the cryo-electron microscopy structures of the human sGCα1β1 heterodimer in different functional states. These structures revealed that the transducer module bridges the nitric oxide sensor module and the catalytic module. Binding of nitric oxide to the β1 haem-nitric oxide and oxygen binding (H-NOX) domain triggers the structural rearrangement of the sensor module and a conformational switch of the transducer module from bending to straightening. The resulting movement of the N termini of the catalytic domains drives structural changes within the catalytic module, which in turn boost the enzymatic activity of sGC. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9884.map.gz emd_9884.map.gz | 3.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9884-v30.xml emd-9884-v30.xml emd-9884.xml emd-9884.xml | 12.2 KB 12.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9884.png emd_9884.png | 65.2 KB | ||
| Filedesc metadata |  emd-9884.cif.gz emd-9884.cif.gz | 5.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9884 http://ftp.pdbj.org/pub/emdb/structures/EMD-9884 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9884 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9884 | HTTPS FTP |
-Related structure data
| Related structure data |  6jt1MC  9883C  9885C  9886C  6jt0C  6jt2C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9884.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9884.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.045 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
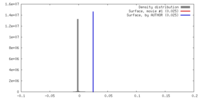
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : human soluble guanylate cyclase
| Entire | Name: human soluble guanylate cyclase |
|---|---|
| Components |
|
-Supramolecule #1: human soluble guanylate cyclase
| Supramolecule | Name: human soluble guanylate cyclase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanylate cyclase soluble subunit alpha-1
| Macromolecule | Name: Guanylate cyclase soluble subunit alpha-1 / type: protein_or_peptide / ID: 1 / Details: GenBank: AAH28384.1 / Number of copies: 1 / Enantiomer: LEVO / EC number: guanylate cyclase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 77.566484 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MFCTKLKDLK ITGECPFSLL APGQVPNESS EEAAGSSESC KATMPICQDI PEKNIQESLP QRKTSRSRVY LHTLAESICK LIFPEFERL NVALQRTLAK HKIKESRKSL EREDFEKTIA EQAVAAGVPV EVIKESLGEE VFKICYEEDE NILGVVGGTL K DFLNSFST ...String: MFCTKLKDLK ITGECPFSLL APGQVPNESS EEAAGSSESC KATMPICQDI PEKNIQESLP QRKTSRSRVY LHTLAESICK LIFPEFERL NVALQRTLAK HKIKESRKSL EREDFEKTIA EQAVAAGVPV EVIKESLGEE VFKICYEEDE NILGVVGGTL K DFLNSFST LLKQSSHCQE AGKRGRLEDA SILCLDKEDD FLHVYYFFPK RTTSLILPGI IKAAAHVLYE TEVEVSLMPP CF HNDCSEF VNQPYLLYSV HMKSTKPSLS PSKPQSSLVI PTSLFCKTFP FHFMFDKDMT ILQFGNGIRR LMNRRDFQGK PNF EEYFEI LTPKINQTFS GIMTMLNMQF VVRVRRWDNS VKKSSRVMDL KGQMIYIVES SAILFLGSPC VDRLEDFTGR GLYL SDIPI HNALRDVVLI GEQARAQDGL KKRLGKLKAT LEQAHQALEE EKKKTVDLLC SIFPCEVAQQ LWQGQVVQAK KFSNV TMLF SDIVGFTAIC SQCSPLQVIT MLNALYTRFD QQCGELDVYK VETIGDAYCV AGGLHKESDT HAVQIALMAV KMMELS DEV MSPHGEPIKM RIGLHSGSVF AGVVGVKMPR YCLFGNNVTL ANKFESCSVP RKINVSPTTY RLLKDCPGFV FTPRSRE EL PPNFPSEIPG ICHFLDAYQQ GTNSKPCFQK KDVEDGNANF LGKASGID UniProtKB: Guanylate cyclase soluble subunit alpha-1 |
-Macromolecule #2: Guanylate cyclase soluble subunit beta-1
| Macromolecule | Name: Guanylate cyclase soluble subunit beta-1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: guanylate cyclase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 70.59932 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MYGFVNHALE LLVIRNYGPE VWEDIKKEAQ LDEEGQFLVR IIYDDSKTYD LVAAASKVLN LNAGEILQMF GKMFFVFCQE SGYDTILRV LGSNVREFLQ NLDALHDHLA TIYPGMRAPS FRCTDAEKGK GLILHYYSER EGLQDIVIGI IKTVAQQIHG T EIDMKVIQ ...String: MYGFVNHALE LLVIRNYGPE VWEDIKKEAQ LDEEGQFLVR IIYDDSKTYD LVAAASKVLN LNAGEILQMF GKMFFVFCQE SGYDTILRV LGSNVREFLQ NLDALHDHLA TIYPGMRAPS FRCTDAEKGK GLILHYYSER EGLQDIVIGI IKTVAQQIHG T EIDMKVIQ QRNEECDHTQ FLIEEKESKE EDFYEDLDRF EENGTQESRI SPYTFCKAFP FHIIFDRDLV VTQCGNAIYR VL PQLQPGN CSLLSVFSLV RPHIDISFHG ILSHINTVFV LRSKEGLLDV EKLECEDELT GTEISCLRLK GQMIYLPEAD SIL FLCSPS VMNLDDLTRR GLYLSDIPLH DATRDLVLLG EQFREEYKLT QELEILTDRL QLTLRALEDE KKKTDTLLYS VLPP SVANE LRHKRPVPAK RYDNVTILFS GIVGFNAFCS KHASGEGAMK IVNLLNDLYT RFDTLTDSRK NPFVYKVETV GDKYM TVSG LPEPCIHHAR SICHLALDMM EIAGQVQVDG ESVQITIGIH TGEVVTGVIG QRMPRYCLFG NTVNLTSRTE TTGEKG KIN VSEYTYRCLM SPENSDPQFH LEHRGPVSMK GKKEPMQVWF LSRKNTGTEE TKQDDD UniProtKB: Guanylate cyclase soluble subunit beta-1 |
-Macromolecule #3: PROTOPORPHYRIN IX CONTAINING FE
| Macromolecule | Name: PROTOPORPHYRIN IX CONTAINING FE / type: ligand / ID: 3 / Number of copies: 1 / Formula: HEM |
|---|---|
| Molecular weight | Theoretical: 616.487 Da |
| Chemical component information |  ChemComp-HEM: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.9 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.0) / Number images used: 379909 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)