+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9831 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of RyR2 (F/A/C/Ca2+ dataset) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cryo-EM / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology information: / negative regulation of calcium-mediated signaling / negative regulation of insulin secretion involved in cellular response to glucose stimulus / neuronal action potential propagation / insulin secretion involved in cellular response to glucose stimulus / negative regulation of release of sequestered calcium ion into cytosol / response to redox state / negative regulation of heart rate / 'de novo' protein folding / FK506 binding ...: / negative regulation of calcium-mediated signaling / negative regulation of insulin secretion involved in cellular response to glucose stimulus / neuronal action potential propagation / insulin secretion involved in cellular response to glucose stimulus / negative regulation of release of sequestered calcium ion into cytosol / response to redox state / negative regulation of heart rate / 'de novo' protein folding / FK506 binding / smooth muscle contraction / T cell proliferation / calcium channel inhibitor activity / Ion homeostasis / regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / release of sequestered calcium ion into cytosol / regulation of cardiac muscle contraction by regulation of the release of sequestered calcium ion / sarcoplasmic reticulum membrane / calcium channel complex / peptidylprolyl isomerase / peptidyl-prolyl cis-trans isomerase activity / calcium channel regulator activity / calcium-mediated signaling / protein maturation / Stimuli-sensing channels / Z disc / protein refolding / positive regulation of cytosolic calcium ion concentration / transmembrane transporter binding / signaling receptor binding / membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
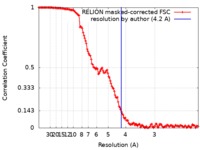
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Gong DS / Chi XM | |||||||||
 Citation Citation |  Journal: Nature / Year: 2019 Journal: Nature / Year: 2019Title: Modulation of cardiac ryanodine receptor 2 by calmodulin. Authors: Deshun Gong / Ximin Chi / Jinhong Wei / Gewei Zhou / Gaoxingyu Huang / Lin Zhang / Ruiwu Wang / Jianlin Lei / S R Wayne Chen / Nieng Yan /    Abstract: The high-conductance intracellular calcium (Ca) channel RyR2 is essential for the coupling of excitation and contraction in cardiac muscle. Among various modulators, calmodulin (CaM) regulates RyR2 ...The high-conductance intracellular calcium (Ca) channel RyR2 is essential for the coupling of excitation and contraction in cardiac muscle. Among various modulators, calmodulin (CaM) regulates RyR2 in a Ca-dependent manner. Here we reveal the regulatory mechanism by which porcine RyR2 is modulated by human CaM through the structural determination of RyR2 under eight conditions. Apo-CaM and Ca-CaM bind to distinct but overlapping sites in an elongated cleft formed by the handle, helical and central domains. The shift in CaM-binding sites on RyR2 is controlled by Ca binding to CaM, rather than to RyR2. Ca-CaM induces rotations and intradomain shifts of individual central domains, resulting in pore closure of the PCB95 and Ca-activated channel. By contrast, the pore of the ATP, caffeine and Ca-activated channel remains open in the presence of Ca-CaM, which suggests that Ca-CaM is one of the many competing modulators of RyR2 gating. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9831.map.gz emd_9831.map.gz | 228.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9831-v30.xml emd-9831-v30.xml emd-9831.xml emd-9831.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_9831_fsc.xml emd_9831_fsc.xml | 14.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_9831.png emd_9831.png | 318.6 KB | ||
| Filedesc metadata |  emd-9831.cif.gz emd-9831.cif.gz | 8.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9831 http://ftp.pdbj.org/pub/emdb/structures/EMD-9831 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9831 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9831 | HTTPS FTP |
-Related structure data
| Related structure data |  6ji0MC  9833C  9834C  9836C  9837C  9879C  9880C  9889C  6ji8C  6jiiC  6jiuC  6jiyC  6jrrC  6jrsC  6jv2C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10241 (Title: afTMEM16/nanodisc complex in the absence of Ca2+ / Data size: 162.0 EMPIAR-10241 (Title: afTMEM16/nanodisc complex in the absence of Ca2+ / Data size: 162.0 Data #1: motion corrected 2D micrographs from 45 frames of afTMEM16/nanodisc complex in the absence of Ca2+ [micrographs - single frame]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9831.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9831.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.091 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : RyR2 in complex with FKBP12.6
| Entire | Name: RyR2 in complex with FKBP12.6 |
|---|---|
| Components |
|
-Supramolecule #1: RyR2 in complex with FKBP12.6
| Supramolecule | Name: RyR2 in complex with FKBP12.6 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: RyR2
| Macromolecule | Name: RyR2 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 564.905625 KDa |
| Sequence | String: MADGGEGEDE IQFLRTDDEV VLQCTATIHK EQQKLCLAAE GFGNRLCFLE STSNSKNVPP DLSICTFVLE QSLSVRALQE MLANTVEKS EGQVDVEKWK FMMKTAQGGG HRTLLYGHAI LLRHSYSGMY LCCLSTSRSS TDKLAFDVGL QEDTTGEACW W TIHPASKQ ...String: MADGGEGEDE IQFLRTDDEV VLQCTATIHK EQQKLCLAAE GFGNRLCFLE STSNSKNVPP DLSICTFVLE QSLSVRALQE MLANTVEKS EGQVDVEKWK FMMKTAQGGG HRTLLYGHAI LLRHSYSGMY LCCLSTSRSS TDKLAFDVGL QEDTTGEACW W TIHPASKQ RSEGEKVRVG DDLILVSVSS ERYLHLSYGN VSLHVDAAFQ QTLWSVAPIS SGSEAAQGYL IGGDVLRLLH GH MDECLTV PSGEHGEEQR RTVHYEGGAV SVHARSLWRL ETLRVAWSGS HIRWGQPFRL RHVTTGKYLS LMEDKSLLLM DKE KADVKS TAFTFRSSKE KLDVGVRKEV DGMGTSEIKY GDSVCFIQHI GTGLWLTYQS VDVKSVRMGS IQRKAIMHHE GHMD DGLNL SRSQHEESRT ARVIRSTVFL FNRFIRGLDA LSKKAKASTV DLPIESVSLS LQDLIGYFHP PDEHLEHEDK QNRLR ALKN RQNLFQEEGM INLVLECIDR LHVYSSAAHF ADVAGREAGE SWKSILNSLY ELLAALIRGN RKNCAQFSGS LDWLIS RLE RLEASSGILE VLHCVLVESP EALNIIKEGH IKSIISLLDK HGRNHKVLDV LCSLCVCHGV AVRSNQHLIC DNLLPGR DL LLQTRLVNHV SSMRPNIFLG VSEGSAQYKK WYYELMVDHT EPFVTAEATH LRVGWASTEG YSPYPGGGEE WGGNGVGD D LFSYGFDGLH LWSGCIARTV SSPNQHLLRT DDVISCCLDL SAPSISFRIN GQPVQGMFEN FNIDGLFFPV VSFSAGIKV RFLLGGRHGE FKFLPPPGYA PCYEAVLPKE KLKVEHSREY KQERTYTRDL LGPTVSLTQA AFTPIPVDTS QIVLPPHLER IREKLAENI HELWVMNKIE LGWQYGPVRD DNKRQHPCLV EFSKLPEQER NYNLQMSLET LKTLLALGCH VGISDEHAEE K VKKMKLPK NYQLTSGYKP APMDLSFIKL TPSQEAMVDK LAENAHNVWA RDRIRQGWTY GIQQDVKNRR NPRLVPYALL DD RTKKSNK DSLREAVRTL LGYGYNLEAP DQDHAARAEV CSGTGERFRI FRAEKTYAVK AGRWYFEFEA VTAGDMRVGW SRP GCQPDQ ELGSDERAFA FDGFKAQRWH QGNEHYGRSW QAGDVVGCMV DMTEHTMMFT LNGEILLDDS GSELAFKDFD VGDG FIPVC SLGVAQVGRM NFGKDVSTLK YFTICGLQEG YEPFAVNTNR DITMWLSKRL PQFLQVPSSH EHIEVTRIDG TIDSS PCLK VTQKSFGSQN SSTDIMFYRL SMPIECAEVF SKTSAGGIPG ASLFGPKNDL EDYDADSDFE VLMKTAHGHL VPDRVD KDK EATKPEFNNH KDYAQEKPSR LKQRFLLRRT KPDYSTSHSA RLTEDVLADD RDDYDYLMQT STYYYSVRIF PGQEPAN VW VGWITSDFHQ YDTAFDLDRV RTVTVTLGDE KGKVHESIKR SNCYMVCAGE SMSPGQGRNN NGLEIGCVVD AASGLLTF T ANGKDLSTYY QVEPSTKLFP AVFAQATSPN VFQFELGRIK NVMPLSAGLF KSEHKNPVPQ CPPRLHVQFL SHVLWSRMP NQFLKVDVSR ISERQGWLVQ CLEPLQFMSL HIPEENRSVD ILELTEQEEL LKFHYHTLRL YSAVCALGNH RVAHALCSHV DEPQLLYAI ENKYMPGLLR AGYYDLLIDI HLSSYATARL MMNNEFIVPM TEETKSITLF PDENKKHGLP GIGLSTSLRP R MQFSSPSF VSINNECYQY SPEFPLDILK AKTIQMLTEA VQEGSLHARD PVGGTTEFLF VPLIKLFYTL LIMGIFHNED LK HILQLIE PSVFKEAAGP EEESDTLEKE PCASEDSRLE GPAEEESKGG KRPKEGLLQM KLPEPVKLQM CLLLQYLCDC QVR HRIEAI VAFSDDFVAK LQDNQRFRYN EVMQALNMSA ALTARKTKEF RSPPQEQINM LLNFKDDKSE CPCPEEIRDQ LLDF HEDLM THCGIELDED GSLDGNSDLT IRGRLLSLVE KVTYLKKKQA EKLVESDSKK SSTLQQLISE TMVRWAQESV IEDPE LVRA MFVLLHRQYD GIGGLVRALP KTYTINGVSV EDTINLLASL GQIRSLLSVR MGKEEEKLMI RGLGDIMNNK VFYQHP NLM RALGMHETVM EVMVNVLGGG ESKEITFPKM VANCCRFLCY FCRISRQNQK AMFDHLSYLL ENSSVGLASP AMRGSTP LD VAAASVMDNN ELALALREPD LEKVVRYLAG CGLQSCQMLV SKGYPDIGWN PVEGERYLDF LRFAVFCNGE SVEENANV V VRLLIRRPEC FGPALRGEGG NGLLAAMEEA IKIAEDPSRD GPSPTSGSSK MPDTEGEEDD TIHMGNAIMT FYAALIDLL GRCAPEMHLI HAAKGEAIRI RSILRSLIPL GDLVGVISIA FQMPTIAKDG NVVEPDMSAG FCPDHKAAMV LFLDRVYGIE VQDFLLHLL EVGFLPDLRA AASLDTAALS ATDMALALNR YLCTAVLPLL TRCAPLFAGT EHHASLIDSL LHTVYRLSKG C SLTKAQRD SIEVCLLSIC GQLRPSMMQH LLRRLVFDVP LLNEHAKMPL KLLTNHYERC WKYYCLPGGW GNFGAASEEE LH LSRKLFW GIFDALSQKK YEQELFKLAL PCLSAVAGAL PPDYMESNYV SMMEKQSSMD SEGNFNPQPV DTSNITIPEK LEY FINKYA EHSHDKWSMD KLANGWIYGE IYSDSSKVQP LMKPYKLLSE KEKEIYRWPI KESLKTMLAW GWRIERTREG DSMA LYNRT RRISQTSQVS VDAAHGYSPR AIDMSNVTLS RDLHAMAEMM AENYHNIWAK KKKLELESKG GGNHPLLVPY DTLTA KEKA KDREKAQDIL KFLQINGYAV SRGFKDLELD TPSIEKRFAY SFLQQLIRYV DEAHQYILEF DGGSRSKGEH FPYEQE IKF FAKVVLPLID QYFKNHRLYF LSAASRPLCS GGHASNKEKE MVTSLFCKLG VLVRHRISLF GNDATSIVNC LHILGQT LD ARTVMKTGLE SVKSALRAFL DNAAEDLEKT MENLKQGQFT HTRNQPKGVT QIINYTTVAL LPMLSSLFEH IGQHQFGE D LILEDVQVSC YRILTSLYAL GTSKSIYVER QRSALGECLA AFAGAFPVAF LETHLDKHNI YSIYNTKSSR ERAALNLPT NVEDVCPNIP SLEKLMEEIV DLAESGIRYT QMPHVMEVVL PMLCSYMSRW WEHGPENNPG RAEMCCTALN SEHMNTLLGN ILKIIYNNL GIDEGAWMKR LAVFSQPIIN KVKPQLLKTH FLPLMEKLKK KAAMVVSEED HLKSEVRGDM SEAELLILDE F TTLARDLY AFYPLLIRFV DYNRAKWLKE PNPEAEDLFR MVAEVFIYWS KSHNFKREEQ NFVVQNEINN MSFLITDTKS KM SKAAVSD QERKKMKRKG DRYSMQTSLI VAALKRLLPI GLNICAPGDQ ELIALAKNRF SLKDTEDEVR DIIRSNIHLQ GKL EDPAIR WQMALYKDLP NRTEDTSDPE KTVERVLDIA NVLFHLEQKS TCMRRRYYSL VEHPQRSKKA VWHKLLSKQR KRAV VACFR MAPLYNLPRH RAVNLFLQGY EKSWIETEEH YFEDKLIEDL AKPGAVPPEE DEGTKRVDPL HQLILLFSRT ALTEK CKLE EDFLYMAYAD IMAKSCHDEE DDDGEEEVKS FEEKEMEKQK LLYQQARLHD RGAAEMVLQT ISASKGETGP MVAATL KLG IAILNGGNST VQQKMLEYLK EKKDVGFFQS LAGLMQSCSV LDLNAFERQN KAEGLGMVTE EGSGEKVLQD DEFTCDL FR FLQLLCEGHN SDFQNYLRTQ TGNNTTVNII ISTVDYLLRV QESISDFYWY YSGKDVIDEQ GQRNFSKAIQ VAKQVFNT L TEYIQGPCTG NQQSLAHSRL WDAVVGFLHV FAHMQMKLSQ DSSQIELLKE LMDLQKDMVV MLLSMLEGNV VNGTIGKQM VDMLVESSNN VEMILKFFDM FLKLKDLTSS DTFKEYDPDG KGVISKRDFH KAMESHKHYT QSETEFLLSC AETDENETLD YEEFVKRFH EPAKDIGFNV AVLLTNLSEH MPNDTRLQTF LELAESVLNY FQPFLGRIEI MGSAKRIERV YFEISESSRT Q WEKPQVKE SKRQFIFDVV NEGGEKEKME LFVNFCEDTI FEMQLAAQIS ESDLNERSAN KEESEKEKPE EQGPRMGFFS LV TVRSALL ALRYNVLTLM RMLSLKSLKK QMKKVKKMTV RDMVTAFFTS YWSVFMTLLH FAASVSRGFS RIIGGLLLGG SLV EGAKKI KVAELLANMP DPTQDEVRGD GDEGERKVLE GTLPSEDLTD LKELTEESDL LSDIFGLDLK REGGQYKLIP HNPN AGLSD LMSSPAPIPE VQEKFQEQKA KEEEKEEKEE NKSEPEKAEG EDGEKEEKAK EDKGKQKLRQ LHTHRYGEPE VPESA FWKK IIAYQQKLLN YFARNFYNMR MLALFVAFAI NFILLFYKVS TSSVVEGKEL PTRSSSENAN FGSLDSSSPR IIAVHY VLE ESSGYMEPTL RILAILHTVI SFFCIIGYYC LKVPLVIFKR EKEVARKLEF DGLYITEQPS EDDIKGQWDR LVINTQS FP NNYWDKFVKR KVMDKYGEFY GRDRISELLG MDKAALDFSD AREKKKPKKD SSLSAVLNSI DVKYQMWKLG VVFTDNSF L YLAWYMTMSV LGHYNNFFFA AHLLDIAMGF KTLRTILSSV THNGKQLVLT VGLLAVVVYL YTVVAFNFFR KFYNKSEDG DTPDMKCDDM LTCYMFHMYV GVRAGGGIGD EIEDPAGDEY EIYRIIFDIT FFFFVIVILL AIIQGLIIDA FGELRDQQEQ VKEDMETKC FICGIGNDYF DTVPHGFETH TLQEHNLANY LFFLMYLINK DETEHTGQES YVWKMYQERC WEFFPAGDCF R KQYEDQLN |
-Macromolecule #2: Peptidyl-prolyl cis-trans isomerase FKBP1B
| Macromolecule | Name: Peptidyl-prolyl cis-trans isomerase FKBP1B / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO / EC number: peptidylprolyl isomerase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.798501 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGVEIETISP GDGRTFPKKG QTCVVHYTGM LQNGKKFDSS RDRNKPFKFR IGKQEVIKGF EEGAAQMSLG QRAKLTCTPD VAYGATGHP GVIPPNATLI FDVELLNLE UniProtKB: Peptidyl-prolyl cis-trans isomerase FKBP1B |
-Macromolecule #3: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 3 / Number of copies: 4 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #4: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 4 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #5: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 5 / Number of copies: 4 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #6: CAFFEINE
| Macromolecule | Name: CAFFEINE / type: ligand / ID: 6 / Number of copies: 4 / Formula: CFF |
|---|---|
| Molecular weight | Theoretical: 194.191 Da |
| Chemical component information |  ChemComp-CFF: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)