登録情報 データベース : EMDB / ID : EMD-9648タイトル Cryo-EM structure of gamma secretase in complex with a Notch fragment 複合体 : gamma-secretaseタンパク質・ペプチド : Nicastrinタンパク質・ペプチド : Presenilin-1タンパク質・ペプチド : Gamma-secretase subunit APH-1Aタンパク質・ペプチド : Gamma-secretase subunit PEN-2タンパク質・ペプチド : Notch1リガンド : 2-acetamido-2-deoxy-beta-D-glucopyranoseリガンド : 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINEリガンド : CHOLESTEROL / 機能・相同性 分子機能 ドメイン・相同性 構成要素
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Homo sapiens (ヒト)手法 / / 解像度 : 2.7 Å Yang G / Zhou R 資金援助 Organization Grant number 国 National Natural Science Foundation of China 31621092
ジャーナル : Nature / 年 : 2019タイトル : Structural basis of Notch recognition by human γ-secretase.著者 : Guanghui Yang / Rui Zhou / Qiang Zhou / Xuefei Guo / Chuangye Yan / Meng Ke / Jianlin Lei / Yigong Shi / 要旨 : Aberrant cleavage of Notch by γ-secretase leads to several types of cancer, but how γ-secretase recognizes its substrate remains unknown. Here we report the cryo-electron microscopy structure of ... Aberrant cleavage of Notch by γ-secretase leads to several types of cancer, but how γ-secretase recognizes its substrate remains unknown. Here we report the cryo-electron microscopy structure of human γ-secretase in complex with a Notch fragment at a resolution of 2.7 Å. The transmembrane helix of Notch is surrounded by three transmembrane domains of PS1, and the carboxyl-terminal β-strand of the Notch fragment forms a β-sheet with two substrate-induced β-strands of PS1 on the intracellular side. Formation of the hybrid β-sheet is essential for substrate cleavage, which occurs at the carboxyl-terminal end of the Notch transmembrane helix. PS1 undergoes pronounced conformational rearrangement upon substrate binding. These features reveal the structural basis of Notch recognition and have implications for the recruitment of the amyloid precursor protein by γ-secretase. 履歴 登録 2018年9月9日 - ヘッダ(付随情報) 公開 2018年12月26日 - マップ公開 2018年12月26日 - 更新 2025年9月17日 - 現状 2025年9月17日 処理サイト : PDBj / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
基本情報 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報 Homo sapiens (ヒト)
Homo sapiens (ヒト) データ登録者
データ登録者 中国, 1件
中国, 1件  引用
引用 ジャーナル: Nature / 年: 2019
ジャーナル: Nature / 年: 2019
 構造の表示
構造の表示 ムービービューア
ムービービューア SurfView
SurfView Molmil
Molmil Jmol/JSmol
Jmol/JSmol ダウンロードとリンク
ダウンロードとリンク emd_9648.map.gz
emd_9648.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-9648-v30.xml
emd-9648-v30.xml emd-9648.xml
emd-9648.xml EMDBヘッダ
EMDBヘッダ emd_9648.png
emd_9648.png emd-9648.cif.gz
emd-9648.cif.gz emd_9648_additional.map.gz
emd_9648_additional.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-9648
http://ftp.pdbj.org/pub/emdb/structures/EMD-9648 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9648
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9648 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
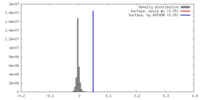
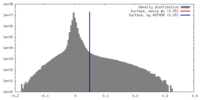
マップ ダウンロード / ファイル: emd_9648.map.gz / 形式: CCP4 / 大きさ: 125 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_9648.map.gz / 形式: CCP4 / 大きさ: 125 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素 Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト)


 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 ムービー
ムービー コントローラー
コントローラー





































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)