[English] 日本語
 Yorodumi
Yorodumi- EMDB-7884: BG505 MD64 N332-GT5 SOSIP trimer in complex with BG18-like precur... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7884 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | BG505 MD64 N332-GT5 SOSIP trimer in complex with BG18-like precursor HMP1 fragment antigen binding and base-binding RM20A3 fragment antigen binding | ||||||||||||
 Map data Map data | Sharpened map | ||||||||||||
 Sample Sample |
| ||||||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell ...symbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / membrane / identical protein binding Similarity search - Function | ||||||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||
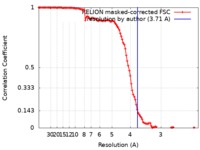
| Method | single particle reconstruction / cryo EM / Resolution: 3.71 Å | ||||||||||||
 Authors Authors | Torres JL / Ozorowski G / Steichen JM / Schief WR / Ward AB | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Science / Year: 2019 Journal: Science / Year: 2019Title: A generalized HIV vaccine design strategy for priming of broadly neutralizing antibody responses. Authors: Jon M Steichen / Ying-Cing Lin / Colin Havenar-Daughton / Simone Pecetta / Gabriel Ozorowski / Jordan R Willis / Laura Toy / Devin Sok / Alessia Liguori / Sven Kratochvil / Jonathan L Torres ...Authors: Jon M Steichen / Ying-Cing Lin / Colin Havenar-Daughton / Simone Pecetta / Gabriel Ozorowski / Jordan R Willis / Laura Toy / Devin Sok / Alessia Liguori / Sven Kratochvil / Jonathan L Torres / Oleksandr Kalyuzhniy / Eleonora Melzi / Daniel W Kulp / Sebastian Raemisch / Xiaozhen Hu / Steffen M Bernard / Erik Georgeson / Nicole Phelps / Yumiko Adachi / Michael Kubitz / Elise Landais / Jeffrey Umotoy / Amanda Robinson / Bryan Briney / Ian A Wilson / Dennis R Burton / Andrew B Ward / Shane Crotty / Facundo D Batista / William R Schief /  Abstract: Vaccine induction of broadly neutralizing antibodies (bnAbs) to HIV remains a major challenge. Germline-targeting immunogens hold promise for initiating the induction of certain bnAb classes; yet for ...Vaccine induction of broadly neutralizing antibodies (bnAbs) to HIV remains a major challenge. Germline-targeting immunogens hold promise for initiating the induction of certain bnAb classes; yet for most bnAbs, a strong dependence on antibody heavy chain complementarity-determining region 3 (HCDR3) is a major barrier. Exploiting ultradeep human antibody sequencing data, we identified a diverse set of potential antibody precursors for a bnAb with dominant HCDR3 contacts. We then developed HIV envelope trimer-based immunogens that primed responses from rare bnAb-precursor B cells in a mouse model and bound a range of potential bnAb-precursor human naïve B cells in ex vivo screens. Our repertoire-guided germline-targeting approach provides a framework for priming the induction of many HIV bnAbs and could be applied to most HCDR3-dominant antibodies from other pathogens. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7884.map.gz emd_7884.map.gz | 117 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7884-v30.xml emd-7884-v30.xml emd-7884.xml emd-7884.xml | 27.1 KB 27.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_7884_fsc.xml emd_7884_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_7884.png emd_7884.png | 50 KB | ||
| Masks |  emd_7884_msk_1.map emd_7884_msk_1.map | 125 MB |  Mask map Mask map | |
| Others |  emd_7884_half_map_1.map.gz emd_7884_half_map_1.map.gz emd_7884_half_map_2.map.gz emd_7884_half_map_2.map.gz | 98.4 MB 98.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7884 http://ftp.pdbj.org/pub/emdb/structures/EMD-7884 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7884 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7884 | HTTPS FTP |
-Related structure data
| Related structure data |  6nf5MC  7875C  7876C  7885C  6dfgC  6dfhC  6nfcC  6oc7C  7886 C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7884.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7884.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.15 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_7884_msk_1.map emd_7884_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_7884_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_7884_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : BG505 MD64 N332-GT5 SOSIP trimer in complex with BG18-like precur...
| Entire | Name: BG505 MD64 N332-GT5 SOSIP trimer in complex with BG18-like precursor HMP1 fragment antigen binding and base-binding RM20A3 fragment antigen binding |
|---|---|
| Components |
|
-Supramolecule #1: BG505 MD64 N332-GT5 SOSIP trimer in complex with BG18-like precur...
| Supramolecule | Name: BG505 MD64 N332-GT5 SOSIP trimer in complex with BG18-like precursor HMP1 fragment antigen binding and base-binding RM20A3 fragment antigen binding type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Molecular weight | Theoretical: 720 KDa |
-Supramolecule #2: SOSIP trimer
| Supramolecule | Name: SOSIP trimer / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant strain: HEK293F Homo sapiens (human) / Recombinant strain: HEK293F |
-Supramolecule #3: HMP1 fragment antigen binding
| Supramolecule | Name: HMP1 fragment antigen binding / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant strain: HEK293F Homo sapiens (human) / Recombinant strain: HEK293F |
-Supramolecule #4: RM20A3 fragment antigen binding
| Supramolecule | Name: RM20A3 fragment antigen binding / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #4-#5 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant strain: HEK293F Homo sapiens (human) / Recombinant strain: HEK293F |
-Macromolecule #1: BG505 MD64 N332-GT5 SOSIP
| Macromolecule | Name: BG505 MD64 N332-GT5 SOSIP / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AENLWVTVYY GVPVWKDAET TLFCASDAKA YETEKHNVWA THACVPTDPN PQEIHLENVT EEFNMWKNNM VEQMHEDIIS LWDQSLKPCV KLTPLCVTLQ CTNYAPKLRS MMRGEIKNCS FNMTTELRDK KQKVYSLFYR LDVVQINENQ GNRSNNSNKE YRLINCNTSA ...String: AENLWVTVYY GVPVWKDAET TLFCASDAKA YETEKHNVWA THACVPTDPN PQEIHLENVT EEFNMWKNNM VEQMHEDIIS LWDQSLKPCV KLTPLCVTLQ CTNYAPKLRS MMRGEIKNCS FNMTTELRDK KQKVYSLFYR LDVVQINENQ GNRSNNSNKE YRLINCNTSA ITQACPKVSF EPIPIHYCAP AGFAILKCKD KKFNGTGPCP SVSTVQCTHG IKPVVSTQLL LNGSLAEEEV IIRSENITNN AKNILVQLNT PVQINCTRPS NNTVKSIRIG PGQAFYYFGD VLGHVRMAHC NISKATWNET LGKVVKQLRK HFGNNTIIRF AQSSGGDLEV TTHSFNCGGE FFYCNTSGLF NSTWISNTSV QGSNSTGSND SLILPCWIKQ IINMWQRIGQ AMYAPPIQGV IRCVSNITGL ILTRDGGSTN STTETFRPGG GDMRDNWRSE LYKYKVVKIE PLGVAPTRCK RRVVGRRRRR RAVGIGAVSL GFLGAAGSTM GAASMTLTVQ ARNLLSGIVQ QQSNLLRAPE PQQHLLKDTH WGIKQLQARV LAVEHYLRDQ QLLGIWGCSG KLICCTNVPW NSSWSNRNLS EIWDNMTWLQ WDKEISNYTQ IIYGLLEESQ NQQEKNEQDL LALDGTKHHH HHH |
-Macromolecule #2: BG18-like precursor HMP1 fragment antigen binding light chain
| Macromolecule | Name: BG18-like precursor HMP1 fragment antigen binding light chain type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: SYELTQPPSV SVSPGQTARI TCSGDALPKQ YAYWYQQKPG QAPVLVIYKD SERPSGIPER FSGSSSGTTV TLTISGVQAE DEADYYCQSA DSSGEVFGGG TKLTVL |
-Macromolecule #3: BG18-like precursor HMP1 fragment antigen binding heavy chain
| Macromolecule | Name: BG18-like precursor HMP1 fragment antigen binding heavy chain type: protein_or_peptide / ID: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLQESGPG LVKPSQTLSL TCTVSGGSIS SGSYYWSWIR QPAGKGLEWI GRIYTSGSTN YNPSLKSRVT ISVDTSKNQF SLKLSSVTAA DTAVYYCARE GFTIFGVVTF SEGYYYGMDV WGKGTTVTVS S |
-Macromolecule #4: RM20A3 fragment antigen binding light chain
| Macromolecule | Name: RM20A3 fragment antigen binding light chain / type: protein_or_peptide / ID: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: ALTQPPSVSG SPGQSVTISC TGTSSDIGSY NYVSWYQQHP GKAPKLMIYD VTQRPSGVSD RFSGSKSGNT ASLTISGLQA DDEADYYCSA YAGRQTFYIF GGGTRLTVLG QPKASPTVTL FPPSSEEL |
-Macromolecule #5: RM20A3 fragment antigen binding heavy chain
| Macromolecule | Name: RM20A3 fragment antigen binding heavy chain / type: protein_or_peptide / ID: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLVETGGG LVQPGGSLKL SCRASGYTFS SFAMSWVRQA PGKGLEWVSL INDRGGLTFY VDSVKGRFTI SRDNSKNTLS LQMHSLRDGD TAVYYCATGG MSSALQSSKY YFDFWGQGAL VTVSS |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5.9 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: Detergent (DDM) added immediately prior to grid preparation | ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Atmosphere: OTHER / Details: Gatan Model 950 Advanced Plasma System | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV / Details: 6 seconds blotting time. | ||||||||||||
| Details | SEC purified complex |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3710 pixel / Digitization - Dimensions - Height: 3838 pixel / Digitization - Sampling interval: 5.0 µm / Number grids imaged: 1 / Number real images: 1796 / Average exposure time: 11.75 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 36000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)