[English] 日本語
 Yorodumi
Yorodumi- EMDB-7835: Cryo-EM structure of the human adenosine A1 receptor-Gi2-protein ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7835 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the human adenosine A1 receptor-Gi2-protein complex bound to its endogenous agonist | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | signaling protein / membrane protein / active-state G protein-coupled receptor / adenosine A1 receptor | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of nucleoside transport / negative regulation of neurotrophin production / regulation of glomerular filtration / negative regulation of circadian sleep/wake cycle, non-REM sleep / G protein-coupled purinergic nucleotide receptor signaling pathway / negative regulation of mucus secretion / purine nucleoside binding / positive regulation of peptide secretion / negative regulation of glutamate secretion / negative regulation of synaptic transmission, GABAergic ...positive regulation of nucleoside transport / negative regulation of neurotrophin production / regulation of glomerular filtration / negative regulation of circadian sleep/wake cycle, non-REM sleep / G protein-coupled purinergic nucleotide receptor signaling pathway / negative regulation of mucus secretion / purine nucleoside binding / positive regulation of peptide secretion / negative regulation of glutamate secretion / negative regulation of synaptic transmission, GABAergic / negative regulation of long-term synaptic depression / positive regulation of dephosphorylation / positive regulation of lipid catabolic process / negative regulation of hormone secretion / mucus secretion / negative regulation of adenylate cyclase-activating adrenergic receptor signaling pathway / negative regulation of leukocyte migration / Muscarinic acetylcholine receptors / G protein-coupled acetylcholine receptor activity / Adenosine P1 receptors / regulation of sensory perception of pain / G protein-coupled adenosine receptor activity / regulation of respiratory gaseous exchange by nervous system process / heterotrimeric G-protein binding / response to purine-containing compound / regulation of presynaptic cytosolic calcium ion concentration / negative regulation of calcium ion-dependent exocytosis / G protein-coupled adenosine receptor signaling pathway / negative regulation of adenylate cyclase activity / positive regulation of potassium ion transport / adenylate cyclase-inhibiting G protein-coupled acetylcholine receptor signaling pathway / positive regulation of neural precursor cell proliferation / negative regulation of systemic arterial blood pressure / negative regulation of synaptic transmission / regulation of cardiac muscle cell contraction / presynaptic active zone / positive regulation of urine volume / negative regulation of synaptic transmission, glutamatergic / long-term synaptic depression / protein targeting to membrane / leukocyte migration / triglyceride homeostasis / gamma-aminobutyric acid signaling pathway / regulation of calcium ion transport / regulation of locomotion / temperature homeostasis / negative regulation of acute inflammatory response / positive regulation of systemic arterial blood pressure / detection of temperature stimulus involved in sensory perception of pain / negative regulation of long-term synaptic potentiation / negative regulation of apoptotic signaling pathway / asymmetric synapse / axolemma / fatty acid homeostasis / phagocytosis / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / negative regulation of lipid catabolic process / neuronal dense core vesicle / lipid catabolic process / positive regulation of vascular associated smooth muscle cell proliferation / positive regulation of superoxide anion generation / heat shock protein binding / Adenylate cyclase inhibitory pathway / response to nutrient / calyx of Held / hippocampal mossy fiber to CA3 synapse / apoptotic signaling pathway / Regulation of insulin secretion / excitatory postsynaptic potential / G protein-coupled receptor binding / vasodilation / cognition / G-protein beta/gamma-subunit complex binding / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / terminal bouton / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / cell-cell signaling / ADP signalling through P2Y purinoceptor 1 Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Draper-Joyce CJ / Khoshouei M | |||||||||
 Citation Citation |  Journal: Nature / Year: 2018 Journal: Nature / Year: 2018Title: Structure of the adenosine-bound human adenosine A receptor-G complex. Authors: Christopher J Draper-Joyce / Maryam Khoshouei / David M Thal / Yi-Lynn Liang / Anh T N Nguyen / Sebastian G B Furness / Hariprasad Venugopal / Jo-Anne Baltos / Jürgen M Plitzko / Radostin ...Authors: Christopher J Draper-Joyce / Maryam Khoshouei / David M Thal / Yi-Lynn Liang / Anh T N Nguyen / Sebastian G B Furness / Hariprasad Venugopal / Jo-Anne Baltos / Jürgen M Plitzko / Radostin Danev / Wolfgang Baumeister / Lauren T May / Denise Wootten / Patrick M Sexton / Alisa Glukhova / Arthur Christopoulos /     Abstract: The class A adenosine A receptor (AR) is a G-protein-coupled receptor that preferentially couples to inhibitory G heterotrimeric G proteins, has been implicated in numerous diseases, yet remains ...The class A adenosine A receptor (AR) is a G-protein-coupled receptor that preferentially couples to inhibitory G heterotrimeric G proteins, has been implicated in numerous diseases, yet remains poorly targeted. Here we report the 3.6 Å structure of the human AR in complex with adenosine and heterotrimeric G protein determined by Volta phase plate cryo-electron microscopy. Compared to inactive AR, there is contraction at the extracellular surface in the orthosteric binding site mediated via movement of transmembrane domains 1 and 2. At the intracellular surface, the G protein engages the AR primarily via amino acids in the C terminus of the Gα α5-helix, concomitant with a 10.5 Å outward movement of the AR transmembrane domain 6. Comparison with the agonist-bound β adrenergic receptor-G-protein complex reveals distinct orientations for each G-protein subtype upon engagement with its receptor. This active AR structure provides molecular insights into receptor and G-protein selectivity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7835.map.gz emd_7835.map.gz | 3.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7835-v30.xml emd-7835-v30.xml emd-7835.xml emd-7835.xml | 15.4 KB 15.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7835.png emd_7835.png | 121.8 KB | ||
| Filedesc metadata |  emd-7835.cif.gz emd-7835.cif.gz | 6.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7835 http://ftp.pdbj.org/pub/emdb/structures/EMD-7835 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7835 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7835 | HTTPS FTP |
-Related structure data
| Related structure data |  6d9hMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7835.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7835.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
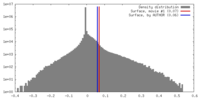
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Human adenosine A1 receptor-Gi2-protein complex bound to its endo...
| Entire | Name: Human adenosine A1 receptor-Gi2-protein complex bound to its endogenous agonist adenosine |
|---|---|
| Components |
|
-Supramolecule #1: Human adenosine A1 receptor-Gi2-protein complex bound to its endo...
| Supramolecule | Name: Human adenosine A1 receptor-Gi2-protein complex bound to its endogenous agonist adenosine type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(i) subunit alpha-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-2 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.502863 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGCTVSAEDK AAAERSKMID KNLREDGEKA AREVKLLLLG AGESGKNTIV KQMKIIHEDG YSEEECRQYR AVVYSNTIQS IMAIVKAMG NLQIDFADPS RADDARQLFA LSCTAEEQGV LPDDLSGVIR RLWADHGVQA CFGRSREYQL NDSAAYYLND L ERIAQSDY ...String: MGCTVSAEDK AAAERSKMID KNLREDGEKA AREVKLLLLG AGESGKNTIV KQMKIIHEDG YSEEECRQYR AVVYSNTIQS IMAIVKAMG NLQIDFADPS RADDARQLFA LSCTAEEQGV LPDDLSGVIR RLWADHGVQA CFGRSREYQL NDSAAYYLND L ERIAQSDY IPTQQDVLRT RVKTTGIVET HFTFKDLHFK MFDVGAQRSE RKKWIHCFEG VTAIIFCVAL SAYDLVLAED EE MNRMHAS MKLFDSICNN KWFTDTSIIL FLNKKDLFEE KITHSPLTIC FPEYTGANKY DEAASYIQSK FEDLNKRKDT KEI YTHFTC STDTKNVQFV FDAVTDVIIK NNLKDCGLF UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-2 |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.534062 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MHHHHHHGSS GSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKL IIWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC R FLDDNQIV ...String: MHHHHHHGSS GSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKL IIWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC R FLDDNQIV TSSGDTTCAL WDIETGQQTT TFTGHTGDVM SLSLAPDTRL FVSGACDASA KLWDVREGMC RQTFTGHESD IN AICFFPN GNAFATGSDD ATCRLFDLRA DQELMTYSHD NIICGITSVS FSKSGRLLLA GYDDFNCNVW DALKADRAGV LAG HDNRVS CLGVTDDGMA VATGSWDSFL KIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: Chimera protein of Muscarinic acetylcholine receptor M4 and Adeno...
| Macromolecule | Name: Chimera protein of Muscarinic acetylcholine receptor M4 and Adenosine receptor A1 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 43.537309 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MKTIIALSYI FCLVFADYKD DDDAMGANFT PVNGSSGNQS VRLVTSSSLE VLFQGPPPSI SAFQAAYIGI EVLIALVSVP GNVLVIWAV KVNQALRDAT FCFIVSLAVA DVAVGALVIP LAILINIGPQ TYFHTCLMVA CPVLILTQSS ILALLAIAVD R YLRVKIPL ...String: MKTIIALSYI FCLVFADYKD DDDAMGANFT PVNGSSGNQS VRLVTSSSLE VLFQGPPPSI SAFQAAYIGI EVLIALVSVP GNVLVIWAV KVNQALRDAT FCFIVSLAVA DVAVGALVIP LAILINIGPQ TYFHTCLMVA CPVLILTQSS ILALLAIAVD R YLRVKIPL RYKMVVTPRR AAVAIAGCWI LSFVVGLTPM FGWNNLSAVE RAWAANGSMG EPVIKCEFEK VISMEYMVYF NF FVWVLPP LLLMVLIYLE VFYLIRKQLN KKVSASSGDP QKYYGKELKI AKSLALILFL FALSWLPLHI LNCITLFCPS CHK PSILTY IAIFLTHGNS AMNPIVYAFR IQKFRVTFLK IWNDHFRCQP APPIDEDLPE ERPDDHHHHH HHH UniProtKB: Muscarinic acetylcholine receptor M4, Adenosine receptor A1 |
-Macromolecule #5: ADENOSINE
| Macromolecule | Name: ADENOSINE / type: ligand / ID: 5 / Number of copies: 1 / Formula: ADN |
|---|---|
| Molecular weight | Theoretical: 267.241 Da |
| Chemical component information | 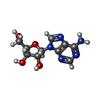 ChemComp-ADN: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Average exposure time: 8.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 47170 / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 3.6 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 263321 |
| Initial angle assignment | Type: COMMON LINE |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller









































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)