[English] 日本語
 Yorodumi
Yorodumi- EMDB-6890: Cryo-EM structure of the human activated spliceosome (late Bact) ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6890 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the human activated spliceosome (late Bact) at 6.5 angstrom | ||||||||||||
 Map data Map data | Cryo-EM structure of the human activated spliceosome (late Bact) at 6.5 angstrom | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | spliceosome / cryo-EM structure / activated spliceosome / late Bact complex / pre-mRNA splicing / SPLICING | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationRES complex / U11/U12 snRNP / post-mRNA release spliceosomal complex / 3'-5' RNA helicase activity / U2 snRNP binding / U7 snRNA binding / histone pre-mRNA DCP binding / generation of catalytic spliceosome for first transesterification step / U7 snRNP / cis assembly of pre-catalytic spliceosome ...RES complex / U11/U12 snRNP / post-mRNA release spliceosomal complex / 3'-5' RNA helicase activity / U2 snRNP binding / U7 snRNA binding / histone pre-mRNA DCP binding / generation of catalytic spliceosome for first transesterification step / U7 snRNP / cis assembly of pre-catalytic spliceosome / histone pre-mRNA 3'end processing complex / SLBP independent Processing of Histone Pre-mRNAs / SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) / B-WICH complex / miRNA processing / alternative mRNA splicing, via spliceosome / protein methylation / poly(A) binding / 7-methylguanosine cap hypermethylation / U12-type spliceosomal complex / embryonic brain development / U1 snRNP binding / U2-type catalytic step 1 spliceosome / methylosome / pre-mRNA binding / regulation of mRNA splicing, via spliceosome / C2H2 zinc finger domain binding / pICln-Sm protein complex / positive regulation of mRNA splicing, via spliceosome / RNA splicing, via transesterification reactions / mRNA 3'-end processing / sno(s)RNA-containing ribonucleoprotein complex / small nuclear ribonucleoprotein complex / SMN-Sm protein complex / spliceosomal tri-snRNP complex / blastocyst formation / splicing factor binding / P granule / snRNP binding / commitment complex / U2-type precatalytic spliceosome / mRNA cis splicing, via spliceosome / oocyte development / telomerase holoenzyme complex / Transport of Mature mRNA derived from an Intron-Containing Transcript / telomerase RNA binding / U2-type catalytic step 2 spliceosome / U2-type prespliceosome assembly / U2-type spliceosomal complex / U1 snRNP / SAGA complex / RNA Polymerase II Transcription Termination / U2 snRNP / U4 snRNP / U2-type prespliceosome / positive regulation of transcription by RNA polymerase III / protein peptidyl-prolyl isomerization / inner cell mass cell proliferation / K63-linked polyubiquitin modification-dependent protein binding / ubiquitin-ubiquitin ligase activity / cyclosporin A binding / precatalytic spliceosome / lipid biosynthetic process / WD40-repeat domain binding / pattern recognition receptor activity / mRNA 3'-splice site recognition / regulation of RNA splicing / mRNA Splicing - Minor Pathway / positive regulation of transcription by RNA polymerase I / spliceosomal complex assembly / spliceosomal tri-snRNP complex assembly / Prp19 complex / U5 snRNP / U5 snRNA binding / pre-mRNA intronic binding / spliceosomal snRNP assembly / blastocyst development / U6 snRNA binding / U2 snRNA binding / protein K63-linked ubiquitination / protein localization to nucleus / regulation of DNA repair / positive regulation of G1/S transition of mitotic cell cycle / Cajal body / U1 snRNA binding / RNA processing / positive regulation of viral genome replication / U4/U6 x U5 tri-snRNP complex / ovarian follicle development / transcription regulator inhibitor activity / spindle assembly / catalytic step 2 spliceosome / transcription-coupled nucleotide-excision repair / mRNA Splicing - Major Pathway / proteasomal protein catabolic process / lipid droplet / Gene and protein expression by JAK-STAT signaling after Interleukin-12 stimulation / RNA splicing / DNA damage checkpoint signaling Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  unidentified adenovirus unidentified adenovirus | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.5 Å | ||||||||||||
 Authors Authors | Zhang X / Yan C | ||||||||||||
| Funding support |  China, 3 items China, 3 items
| ||||||||||||
 Citation Citation |  Journal: Cell Res / Year: 2018 Journal: Cell Res / Year: 2018Title: Structure of the human activated spliceosome in three conformational states. Authors: Xiaofeng Zhang / Chuangye Yan / Xiechao Zhan / Lijia Li / Jianlin Lei / Yigong Shi /  Abstract: During each cycle of pre-mRNA splicing, the pre-catalytic spliceosome (B complex) is converted into the activated spliceosome (B complex), which has a well-formed active site but cannot proceed to ...During each cycle of pre-mRNA splicing, the pre-catalytic spliceosome (B complex) is converted into the activated spliceosome (B complex), which has a well-formed active site but cannot proceed to the branching reaction. Here, we present the cryo-EM structure of the human B complex in three distinct conformational states. The EM map allows atomic modeling of nearly all protein components of the U2 small nuclear ribonucleoprotein (snRNP), including three of the SF3a complex and seven of the SF3b complex. The structure of the human B complex contains 52 proteins, U2, U5, and U6 small nuclear RNA (snRNA), and a pre-mRNA. Three distinct conformations have been captured, representing the early, mature, and late states of the human B complex. These complexes differ in the orientation of the Switch loop of Prp8, the splicing factors RNF113A and NY-CO-10, and most components of the NineTeen complex (NTC) and the NTC-related complex. Analysis of these three complexes and comparison with the B and C complexes reveal an ordered flux of components in the B-to-B and the B-to-B transitions, which ultimately prime the active site for the branching reaction. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6890.map.gz emd_6890.map.gz | 228.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6890-v30.xml emd-6890-v30.xml emd-6890.xml emd-6890.xml | 78.8 KB 78.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6890.png emd_6890.png | 171.8 KB | ||
| Filedesc metadata |  emd-6890.cif.gz emd-6890.cif.gz | 24.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6890 http://ftp.pdbj.org/pub/emdb/structures/EMD-6890 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6890 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6890 | HTTPS FTP |
-Related structure data
| Related structure data |  5z57MC  6889C  6891C  5z56C  5z58C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6890.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6890.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of the human activated spliceosome (late Bact) at 6.5 angstrom | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.338 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
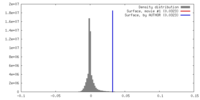
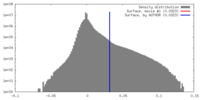
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : late Bact splieosome
+Supramolecule #1: late Bact splieosome
+Macromolecule #1: Pre-mRNA-processing-splicing factor 8
+Macromolecule #3: 116 kDa U5 small nuclear ribonucleoprotein component
+Macromolecule #4: U5 small nuclear ribonucleoprotein 200 kDa helicase
+Macromolecule #5: U5 small nuclear ribonucleoprotein 40 kDa protein
+Macromolecule #6: Small nuclear ribonucleoprotein Sm D3
+Macromolecule #7: Small nuclear ribonucleoprotein-associated proteins B and B'
+Macromolecule #8: Small nuclear ribonucleoprotein Sm D1
+Macromolecule #9: Small nuclear ribonucleoprotein Sm D2
+Macromolecule #10: Small nuclear ribonucleoprotein F
+Macromolecule #11: Small nuclear ribonucleoprotein E
+Macromolecule #12: Small nuclear ribonucleoprotein G
+Macromolecule #16: U2 small nuclear ribonucleoprotein A'
+Macromolecule #17: U2 small nuclear ribonucleoprotein B''
+Macromolecule #18: Splicing factor 3A subunit 3
+Macromolecule #19: Splicing factor 3A subunit 1
+Macromolecule #20: Splicing factor 3A subunit 2
+Macromolecule #21: Splicing factor 3B subunit 1
+Macromolecule #22: Splicing factor 3B subunit 2
+Macromolecule #23: Splicing factor 3B subunit 3
+Macromolecule #24: Splicing factor 3B subunit 4
+Macromolecule #25: Splicing factor 3B subunit 6
+Macromolecule #26: PHD finger-like domain-containing protein 5A
+Macromolecule #27: Splicing factor 3B subunit 5
+Macromolecule #28: Crooked neck-like protein 1
+Macromolecule #29: Cell division cycle 5-like protein
+Macromolecule #30: Pre-mRNA-processing factor 19
+Macromolecule #31: Pre-mRNA-splicing factor SPF27
+Macromolecule #32: Pre-mRNA-splicing factor SYF1
+Macromolecule #33: Intron-binding protein aquarius
+Macromolecule #34: Protein BUD31 homolog
+Macromolecule #35: Pre-mRNA-splicing factor RBM22
+Macromolecule #36: Spliceosome-associated protein CWC15 homolog
+Macromolecule #37: Skip
+Macromolecule #38: Peptidyl-prolyl cis-trans isomerase-like 1
+Macromolecule #39: Pleiotropic regulator 1
+Macromolecule #40: Serine/arginine repetitive matrix protein 2
+Macromolecule #41: Pre-mRNA-splicing factor CWC22 homolog
+Macromolecule #42: Pre-mRNA-processing factor 17
+Macromolecule #43: Smad nuclear-interacting protein 1
+Macromolecule #44: RNA-binding motif protein, X-linked 2
+Macromolecule #45: BUD13 homolog
+Macromolecule #46: Pre-mRNA-splicing factor ATP-dependent RNA helicase DHX16
+Macromolecule #47: Peptidyl-prolyl cis-trans isomerase E
+Macromolecule #2: U5 snRNA
+Macromolecule #13: U6 snRNA
+Macromolecule #14: pre-mRNA
+Macromolecule #15: U2 snRNA
+Macromolecule #48: INOSITOL HEXAKISPHOSPHATE
+Macromolecule #49: ALANINE
+Macromolecule #50: GUANOSINE-5'-TRIPHOSPHATE
+Macromolecule #51: MAGNESIUM ION
+Macromolecule #52: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 / Details: 20 mM HEPES-KOH, pH 7.9, 150 mM NaCl, 1.5 mM MgCl2 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 48.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)