+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5235 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the Drosophila apoptosome | |||||||||
 Map data Map data | map of the Drosophila apoptosome | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Drosophila apoptosome / apoptosis / programmed cell death | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of humoral immune response / positive regulation of glial cell apoptotic process / Formation of apoptosome / salivary gland histolysis / positive regulation of compound eye retinal cell programmed cell death / melanization defense response / sarcosine catabolic process / Activation of caspases through apoptosome-mediated cleavage / Regulation of the apoptosome activity / central nervous system formation ...negative regulation of humoral immune response / positive regulation of glial cell apoptotic process / Formation of apoptosome / salivary gland histolysis / positive regulation of compound eye retinal cell programmed cell death / melanization defense response / sarcosine catabolic process / Activation of caspases through apoptosome-mediated cleavage / Regulation of the apoptosome activity / central nervous system formation / chaeta development / sperm individualization / autophagic cell death / apoptosome / Neutrophil degranulation / CARD domain binding / S-adenosylmethionine cycle / programmed cell death / triglyceride homeostasis / dendrite morphogenesis / response to starvation / cysteine-type endopeptidase activator activity involved in apoptotic process / response to gamma radiation / ADP binding / neuron cellular homeostasis / positive regulation of apoptotic process / ATP binding / identical protein binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.9 Å | |||||||||
 Authors Authors | Yuan S / Yu X / Topf M / Ludtke SJ / Akey CW | |||||||||
 Citation Citation |  Journal: Structure / Year: 2011 Journal: Structure / Year: 2011Title: Structure of the Drosophila apoptosome at 6.9 å resolution. Authors: Shujun Yuan / Xinchao Yu / Maya Topf / Loretta Dorstyn / Sharad Kumar / Steven J Ludtke / Christopher W Akey /  Abstract: The Drosophila Apaf-1 related killer forms an apoptosome in the intrinsic cell death pathway. In this study we show that Dark forms a single ring when initiator procaspases are bound. This Dark-Dronc ...The Drosophila Apaf-1 related killer forms an apoptosome in the intrinsic cell death pathway. In this study we show that Dark forms a single ring when initiator procaspases are bound. This Dark-Dronc complex cleaves DrICE efficiently; hence, a single ring represents the Drosophila apoptosome. We then determined the 3D structure of a double ring at ∼6.9 Å resolution and created a model of the apoptosome. Subunit interactions in the Dark complex are similar to those in Apaf-1 and CED-4 apoptosomes, but there are significant differences. In particular, Dark has "lost" a loop in the nucleotide-binding pocket, which opens a path for possible dATP exchange in the apoptosome. In addition, caspase recruitment domains (CARDs) form a crown on the central hub of the Dark apoptosome. This CARD geometry suggests that conformational changes will be required to form active Dark-Dronc complexes. When taken together, these data provide insights into apoptosome structure, function, and evolution. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5235.map.gz emd_5235.map.gz | 83.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5235-v30.xml emd-5235-v30.xml emd-5235.xml emd-5235.xml | 11.2 KB 11.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5235_1.jpg emd_5235_1.jpg | 208.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5235 http://ftp.pdbj.org/pub/emdb/structures/EMD-5235 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5235 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5235 | HTTPS FTP |
-Validation report
| Summary document |  emd_5235_validation.pdf.gz emd_5235_validation.pdf.gz | 331.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5235_full_validation.pdf.gz emd_5235_full_validation.pdf.gz | 331.4 KB | Display | |
| Data in XML |  emd_5235_validation.xml.gz emd_5235_validation.xml.gz | 6.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5235 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5235 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5235 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5235 | HTTPS FTP |
-Related structure data
| Related structure data |  4v4lMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5235.map.gz / Format: CCP4 / Size: 89 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5235.map.gz / Format: CCP4 / Size: 89 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map of the Drosophila apoptosome | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.72 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
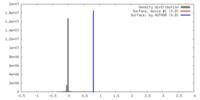
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Drosophila apoptosome (double ring)
| Entire | Name: Drosophila apoptosome (double ring) |
|---|---|
| Components |
|
-Supramolecule #1000: Drosophila apoptosome (double ring)
| Supramolecule | Name: Drosophila apoptosome (double ring) / type: sample / ID: 1000 Details: Sample assembled in low salt buffer (20 mM HEPES pH 7.5, 10 mM KCl, 1.5 mM MgCl2, 1 mM EDTA, 1 mM EGTA, 1 mM DTT) at about 0.5 mg per ml with dATP Oligomeric state: hexadecamer of Dark molecules / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 2.5 MDa |
-Macromolecule #1: Drosophila Apaf-1 like protein
| Macromolecule | Name: Drosophila Apaf-1 like protein / type: protein_or_peptide / ID: 1 / Name.synonym: Dark / Number of copies: 16 / Oligomeric state: hexadecamer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 160 KDa |
| Recombinant expression | Organism: sf21 insect cells (unknown) / Recombinant plasmid: pFastBac |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 20mM HEPES, 10mM KCl, 1.5mM MgCl2, 1mM EDTA, 1mM EGTA, 1mM DTT |
| Grid | Details: thin carbon film covered holey grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 77 K / Instrument: OTHER Details: Vitrification instrument: FEI Vitrobot. blotting at room temperature with sample at room temperature Method: Blot for 2-2.5s before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Temperature | Min: 93 K / Max: 100 K / Average: 93 K |
| Alignment procedure | Legacy - Astigmatism: objective lens astigmatism was corrected at 200,000 times magnification |
| Details | actual magnification at the ccd 87000, camera pixel size 15um, 1.72 angstrom per pixel, data collected semi-automatically with EMTools |
| Date | Sep 15, 2009 |
| Image recording | Category: CCD / Film or detector model: GENERIC TVIPS (4k x 4k) / Number real images: 1100 / Average electron dose: 20 e/Å2 Details: the frames were 4096 x 4096 tiffs collected with the TVIPS CCD |
| Electron beam | Acceleration voltage: 160 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 50000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder: Side entry liquid nitrogen-cooled cryo specimen holder Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | actual class number 1353 |
|---|---|
| CTF correction | Details: each image |
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 6.9 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: EMAN2 Details: Projection matching was done with Fourier ring correlation, model-based masking and SSNR weighting over an 80-6 angstrom resolution range. The final refinement steps used an angular step of ...Details: Projection matching was done with Fourier ring correlation, model-based masking and SSNR weighting over an 80-6 angstrom resolution range. The final refinement steps used an angular step of 2.5 degrees and each of the 48,000 particles was matched to the best two projection classes (1353). In total, 45,000 particles were used in the final reconstruction and the 3D map was amplitude corrected then Gaussian low-pass filtered with a Fourier half-width of 0.12. Number images used: 48271 |
| Final two d classification | Number classes: 1000 |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)