[English] 日本語
 Yorodumi
Yorodumi- EMDB-44260: Cryo-EM structure of the avian great tit TRPM8 channel in complex... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the avian great tit TRPM8 channel in complex with the antagonist TC-I 2014 | |||||||||
 Map data Map data | The final full map sharpened with a B factor of -60 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRPM8 / menthol receptor / cold receptor / PI(4 / 5)P2 / cooling agonists / temperature sensing / ion channel / sensory transduction / transient receptor potential ion channel / TRPM8 activation / TRPM8 inhibition / TRPM8 desensitization / TRPM8 antagonists / MEMBRANE PROTEIN | |||||||||
| Biological species |  Parus major (Great Tit) Parus major (Great Tit) | |||||||||
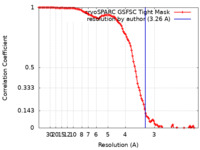
| Method | single particle reconstruction / cryo EM / Resolution: 3.26 Å | |||||||||
 Authors Authors | Yin Y / Park C-G / Zhang F / Fedor J / Feng S / Suo Y / Im W / Lee S-Y | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2024 Journal: Sci Adv / Year: 2024Title: Mechanisms of sensory adaptation and inhibition of the cold and menthol receptor TRPM8. Authors: Ying Yin / Cheon-Gyu Park / Feng Zhang / Justin G Fedor / Shasha Feng / Yang Suo / Wonpil Im / Seok-Yong Lee /  Abstract: Our sensory adaptation to cold and chemically induced coolness is mediated by the intrinsic property of TRPM8 channels to desensitize. TRPM8 is also implicated in cold-evoked pain disorders and ...Our sensory adaptation to cold and chemically induced coolness is mediated by the intrinsic property of TRPM8 channels to desensitize. TRPM8 is also implicated in cold-evoked pain disorders and migraine, highlighting its inhibitors as an avenue for pain relief. Despite the importance, the mechanisms of TRPM8 desensitization and inhibition remained unclear. We found, using cryo-electron microscopy, electrophysiology, and molecular dynamics simulations, that TRPM8 inhibitors bind selectively to the desensitized state of the channel. These inhibitors were used to reveal the overlapping mechanisms of desensitization and inhibition and that cold and cooling agonists share a common desensitization pathway. Furthermore, we identified the structural determinants crucial for the conformational change in TRPM8 desensitization. Our study illustrates how receptor-level conformational changes alter cold sensation, providing insights into therapeutic development. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_44260.map.gz emd_44260.map.gz | 64.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-44260-v30.xml emd-44260-v30.xml emd-44260.xml emd-44260.xml | 20.2 KB 20.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_44260_fsc.xml emd_44260_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_44260.png emd_44260.png | 133.5 KB | ||
| Filedesc metadata |  emd-44260.cif.gz emd-44260.cif.gz | 7.1 KB | ||
| Others |  emd_44260_half_map_1.map.gz emd_44260_half_map_1.map.gz emd_44260_half_map_2.map.gz emd_44260_half_map_2.map.gz | 115.9 MB 115.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-44260 http://ftp.pdbj.org/pub/emdb/structures/EMD-44260 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44260 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44260 | HTTPS FTP |
-Related structure data
| Related structure data |  9b6iMC  9b6dC  9b6eC  9b6fC  9b6gC  9b6hC  9b6jC  9b6kC M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_44260.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_44260.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The final full map sharpened with a B factor of -60 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map A of the final 3D reconstruction
| File | emd_44260_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A of the final 3D reconstruction | ||||||||||||
| Projections & Slices |
| ||||||||||||
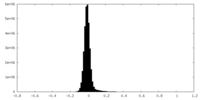
| Density Histograms |
-Half map: Half map B of the final 3D reconstruction
| File | emd_44260_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B of the final 3D reconstruction | ||||||||||||
| Projections & Slices |
| ||||||||||||
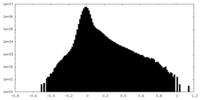
| Density Histograms |
- Sample components
Sample components
-Entire : Transient receptor potential cation channel subfamily M member 8
| Entire | Name: Transient receptor potential cation channel subfamily M member 8 |
|---|---|
| Components |
|
-Supramolecule #1: Transient receptor potential cation channel subfamily M member 8
| Supramolecule | Name: Transient receptor potential cation channel subfamily M member 8 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Parus major (Great Tit) Parus major (Great Tit) |
-Macromolecule #1: Transient receptor potential cation channel subfamily M member 8
| Macromolecule | Name: Transient receptor potential cation channel subfamily M member 8 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Parus major (Great Tit) Parus major (Great Tit) |
| Molecular weight | Theoretical: 131.966203 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAIAEGVLGH ILETLVSMGS MRHRRNGNFE SSRLLYSSMS RSIDVACSDA DLANFIQENF KKRECVFFTK DTKSMGNLCK CGYPENQHI EGTQVNTTEK WNYKKHTKEL PTDAFGDIQF ENLGKRGKYI RLSCDTDSET LYDLMTQHWH LKTPNLVISV T GGAKNFAL ...String: MAIAEGVLGH ILETLVSMGS MRHRRNGNFE SSRLLYSSMS RSIDVACSDA DLANFIQENF KKRECVFFTK DTKSMGNLCK CGYPENQHI EGTQVNTTEK WNYKKHTKEL PTDAFGDIQF ENLGKRGKYI RLSCDTDSET LYDLMTQHWH LKTPNLVISV T GGAKNFAL KPRMRKIFSR LIYIAQSKGA WIFTGGTHYG LMKYIGEVVR DNTISRSSEE NVVAIGIAAW GMISNRETLI RT ADSDGSF LARYIMDDLK RDPLYCLDNN HTHLLLVDNG THGHPTTEAK VRTQLEKYIS ERVIPESNYG GKIPIVCFAQ GGG KETLKS INVAIKSKIP CVVVEGSGRI ADVIASLVEA EGTLASSCVK ESLLRFLPRT ISRLSEEETE SWIKWIKEVL ESPH LLTVI KIEEAGDEIV SNAISFALYK AFSTNEHDRD NWNGQLKLLL EWNQLDLASD EIFTNDRNWE SADLQDVMFT ALVKD RPKF VRLFLENGLN LRKFLTTEVL RELYTNNFSS LVFKNLQIAK NSYNDALLTF VWKMVEDFRR GFKRDYKNSK DEMEIQ LSE ECPITRHPLQ ALFIWSVLQN KKELSKVIWE QTRGCTLAAL GASKLLKSMA KVKNDINAAG ESEELANEYE TRAVELF TE CYSNDEDLAE QLLTYSCEAW GGSNCLELAV EARDQQFIAQ PGVQNFLSKQ WYGEISRDTK NWKIIMCLFF FPLIGCGF I SFRKKPVEKS KKLFLYYVSF FTSPFVVFSW NVIFYIAFLL LFAYVLLMDF QKEPTALEII LYVLVFVLLC DEWYMNGSK YFSDLWNVMD TLAIFYFIAG IVFRLHSDES SWYSGRVIFC LDYIVFTLRL IHIFTVSRNL GPKIIMLQRM MIDVFFFLFL FAVWMVAFG VARQGILRKN EHRWEWIFRS VIYEPYLAMF GQYPDDIDGT TYNFDRCTFS GNESKPLCVE LDANNQPRFP E WITIPLVC IYMLSTNILL VNLLVAMFGY TVGSVQENND QVWKFQRFFL VQEYCSRLTI PFPFVIFAYI FMVMRKCFKC CC NKESKEP SICCSRNEDN EILAWEAVMK ENYLVKINTK ANDSSEEMVH RFRQLDAKLS DLKGLLKEIS SKIKSNSLEV LFQ GPDYKD DDDKAHHHHH HHHHH |
-Macromolecule #2: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #3: CHOLESTEROL HEMISUCCINATE
| Macromolecule | Name: CHOLESTEROL HEMISUCCINATE / type: ligand / ID: 3 / Number of copies: 16 / Formula: Y01 |
|---|---|
| Molecular weight | Theoretical: 486.726 Da |
| Chemical component information |  ChemComp-Y01: |
-Macromolecule #4: 3-{7-(trifluoromethyl)-5-[2-(trifluoromethyl)phenyl]-1H-benzimida...
| Macromolecule | Name: 3-{7-(trifluoromethyl)-5-[2-(trifluoromethyl)phenyl]-1H-benzimidazol-2-yl}-1-oxa-2-azaspiro[4.5]dec-2-ene type: ligand / ID: 4 / Number of copies: 4 / Formula: T14 |
|---|---|
| Molecular weight | Theoretical: 467.407 Da |
| Chemical component information | 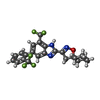 ChemComp-T14: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Software | Name: Latitude |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number real images: 10266 / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Software | Name:  Coot Coot | |||||||||
| Refinement | Space: REAL / Protocol: RIGID BODY FIT | |||||||||
| Output model |  PDB-9b6i: |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)