[English] 日本語
 Yorodumi
Yorodumi- EMDB-43021: 60S ribosome biogenesis intermediate (Dbp10 catalytic structure -... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | 60S ribosome biogenesis intermediate (Dbp10 catalytic structure - Overall map) | ||||||||||||
 Map data Map data | Overall map Dbp10 catalytic intermediate | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Ribosome biogenesis / intermediate DEAD-box ATPase / Ribosome-RNA complex | ||||||||||||
| Function / homology |  Function and homology information Function and homology information25S rRNA (cytosine2870-C5)-methyltransferase / snoRNA release from pre-rRNA / nuclear division / exonucleolytic trimming to generate mature 5'-end of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / endonucleolytic cleavage in ITS1 upstream of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / rRNA (cytosine-C5-)-methyltransferase activity / endonucleolytic cleavage in 5'-ETS of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / endonucleolytic cleavage to generate mature 5'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / PeBoW complex / intracellular mRNA localization ...25S rRNA (cytosine2870-C5)-methyltransferase / snoRNA release from pre-rRNA / nuclear division / exonucleolytic trimming to generate mature 5'-end of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / endonucleolytic cleavage in ITS1 upstream of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / rRNA (cytosine-C5-)-methyltransferase activity / endonucleolytic cleavage in 5'-ETS of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / endonucleolytic cleavage to generate mature 5'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / PeBoW complex / intracellular mRNA localization / rRNA base methylation / rRNA primary transcript binding / ATP-dependent activity, acting on RNA / positive regulation of ATP-dependent activity / maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / rRNA methylation / protein folding chaperone complex / maturation of 5.8S rRNA / hexon binding / preribosome, small subunit precursor / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / regulation of cell size / proteasome binding / Major pathway of rRNA processing in the nucleolus and cytosol / SRP-dependent cotranslational protein targeting to membrane / GTP hydrolysis and joining of the 60S ribosomal subunit / ribosomal large subunit binding / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Formation of a pool of free 40S subunits / preribosome, large subunit precursor / nuclear-transcribed mRNA catabolic process / ATPase activator activity / L13a-mediated translational silencing of Ceruloplasmin expression / translational elongation / ribosomal large subunit export from nucleus / 90S preribosome / mRNA transport / ribonucleoprotein complex binding / ribosomal subunit export from nucleus / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of LSU-rRNA / translation initiation factor activity / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / proteasome complex / nuclear periphery / assembly of large subunit precursor of preribosome / ribosomal large subunit biogenesis / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of SSU-rRNA / cytosolic ribosome assembly / small-subunit processome / macroautophagy / maintenance of translational fidelity / protein catabolic process / rRNA processing / ribosomal small subunit biogenesis / viral capsid / nuclear envelope / ATPase binding / protein-macromolecule adaptor activity / 5S rRNA binding / large ribosomal subunit rRNA binding / ribosomal large subunit assembly / cytoplasmic translation / cytosolic large ribosomal subunit / RNA helicase activity / negative regulation of translation / rRNA binding / RNA helicase / ribosome / structural constituent of ribosome / translation / GTPase activity / mRNA binding / host cell nucleus / GTP binding / nucleolus / ATP hydrolysis activity / DNA binding / RNA binding / nucleoplasm / ATP binding / identical protein binding / nucleus / metal ion binding / cytosol / cytoplasm Similarity search - Function | ||||||||||||
| Biological species |   | ||||||||||||
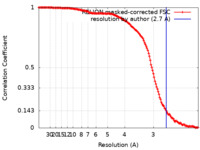
| Method | single particle reconstruction / cryo EM / Resolution: 2.7 Å | ||||||||||||
 Authors Authors | Cruz VE / Weirich CS / Peddada N / Erzberger JP | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: The DEAD-box ATPase Dbp10/DDX54 initiates peptidyl transferase center formation during 60S ribosome biogenesis. Authors: Victor E Cruz / Christine S Weirich / Nagesh Peddada / Jan P Erzberger /  Abstract: DEAD-box ATPases play crucial roles in guiding rRNA restructuring events during the biogenesis of large (60S) ribosomal subunits, but their precise molecular functions are currently unknown. In this ...DEAD-box ATPases play crucial roles in guiding rRNA restructuring events during the biogenesis of large (60S) ribosomal subunits, but their precise molecular functions are currently unknown. In this study, we present cryo-EM reconstructions of nucleolar pre-60S intermediates that reveal an unexpected, alternate secondary structure within the nascent peptidyl-transferase-center (PTC). Our analysis of three sequential nucleolar pre-60S intermediates reveals that the DEAD-box ATPase Dbp10/DDX54 remodels this alternate base pairing and enables the formation of the rRNA junction that anchors the mature form of the universally conserved PTC A-loop. Post-catalysis, Dbp10 captures rRNA helix H61, initiating the concerted exchange of biogenesis factors during late nucleolar 60S maturation. Our findings show that Dbp10 activity is essential for the formation of the ribosome active site and reveal how this function is integrated with subsequent assembly steps to drive the biogenesis of the large ribosomal subunit. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43021.map.gz emd_43021.map.gz | 229.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43021-v30.xml emd-43021-v30.xml emd-43021.xml emd-43021.xml | 76.5 KB 76.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43021_fsc.xml emd_43021_fsc.xml | 14.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_43021.png emd_43021.png | 105.7 KB | ||
| Filedesc metadata |  emd-43021.cif.gz emd-43021.cif.gz | 18.5 KB | ||
| Others |  emd_43021_half_map_1.map.gz emd_43021_half_map_1.map.gz emd_43021_half_map_2.map.gz emd_43021_half_map_2.map.gz | 194.9 MB 194.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43021 http://ftp.pdbj.org/pub/emdb/structures/EMD-43021 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43021 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43021 | HTTPS FTP |
-Validation report
| Summary document |  emd_43021_validation.pdf.gz emd_43021_validation.pdf.gz | 1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_43021_full_validation.pdf.gz emd_43021_full_validation.pdf.gz | 1 MB | Display | |
| Data in XML |  emd_43021_validation.xml.gz emd_43021_validation.xml.gz | 21.6 KB | Display | |
| Data in CIF |  emd_43021_validation.cif.gz emd_43021_validation.cif.gz | 28.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43021 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43021 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43021 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43021 | HTTPS FTP |
-Related structure data
| Related structure data |  8v84MC  8v83C  8v85C  8v87C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_43021.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43021.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Overall map Dbp10 catalytic intermediate | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Overall half-map 1 Dbp10 catalytic intermediate
| File | emd_43021_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Overall half-map 1 Dbp10 catalytic intermediate | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Overall half-map 2 Dbp10 catalytic intermediate
| File | emd_43021_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Overall half-map 2 Dbp10 catalytic intermediate | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : 60S ribosome biogenesis intermediate
+Supramolecule #1: 60S ribosome biogenesis intermediate
+Macromolecule #1: Saccharomyces cerevisiae 25S ribosomal RNA
+Macromolecule #2: Saccharomyces cerevisiae 5.8S ribosomal RNA
+Macromolecule #4: ITS2 RNA
+Macromolecule #3: ATP-dependent RNA helicase DBP10
+Macromolecule #5: 60S ribosomal subunit assembly/export protein LOC1
+Macromolecule #6: Ribosomal RNA-processing protein 14
+Macromolecule #7: Ribosome biogenesis protein BRX1
+Macromolecule #8: Large ribosomal subunit protein uL3
+Macromolecule #9: 60S ribosomal protein L4-A
+Macromolecule #10: ATP-dependent RNA helicase HAS1
+Macromolecule #11: 60S ribosomal protein L6-A
+Macromolecule #12: 60S ribosomal protein L7-A
+Macromolecule #13: Large ribosomal subunit protein eL8A
+Macromolecule #14: Large ribosomal subunit protein uL6A
+Macromolecule #15: Ribosome biogenesis protein NSA1
+Macromolecule #16: rRNA-processing protein EBP2
+Macromolecule #17: Proteasome-interacting protein CIC1
+Macromolecule #18: 60S ribosomal protein L13-A
+Macromolecule #19: 60S ribosomal protein L14-A
+Macromolecule #20: Large ribosomal subunit protein eL15A
+Macromolecule #21: Large ribosomal subunit protein uL13A
+Macromolecule #22: 60S ribosomal protein L17-A
+Macromolecule #23: 60S ribosomal protein L18-A
+Macromolecule #24: Protein MAK16
+Macromolecule #25: Large ribosomal subunit protein eL20A
+Macromolecule #26: Ribosomal RNA-processing protein 15
+Macromolecule #27: 60S ribosomal protein L23-A
+Macromolecule #28: Ribosome assembly factor MRT4
+Macromolecule #29: 60S ribosomal protein L26-A
+Macromolecule #30: Ribosome biogenesis protein SSF1
+Macromolecule #31: Large ribosomal subunit protein uL1A
+Macromolecule #32: Nucleolar GTP-binding protein 1
+Macromolecule #33: 60S ribosomal protein L32
+Macromolecule #34: 60S ribosomal protein L33-A
+Macromolecule #35: 60S ribosomal protein L35-A
+Macromolecule #36: 60S ribosomal protein L36-A
+Macromolecule #37: 60S ribosomal protein L37-A
+Macromolecule #38: 60S ribosome subunit biogenesis protein NIP7
+Macromolecule #39: Ribosome biogenesis protein ERB1
+Macromolecule #40: Pescadillo homolog
+Macromolecule #41: Ribosome biogenesis protein 15
+Macromolecule #42: 25S rRNA (cytosine(2870)-C(5))-methyltransferase
+Macromolecule #43: Ribosome biogenesis protein NSA2
+Macromolecule #44: Ribosome biogenesis protein RLP7
+Macromolecule #45: Ribosome biogenesis protein RLP24
+Macromolecule #46: Nucleolar protein 16
+Macromolecule #47: Ribosomal RNA-processing protein 1
+Macromolecule #48: Ribosome production factor 1
+Macromolecule #49: Eukaryotic translation initiation factor 6
+Macromolecule #50: GUANOSINE-5'-DIPHOSPHATE
+Macromolecule #51: MAGNESIUM ION
+Macromolecule #52: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 39.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.9 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)