[English] 日本語
 Yorodumi
Yorodumi- EMDB-42544: KIF1A[1-393] AMP-PNP bound two-heads-bound state in complex with ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | KIF1A[1-393] AMP-PNP bound two-heads-bound state in complex with a microtubule - class T2L1 | |||||||||||||||||||||
 Map data Map data | Primary map, locally refined on the two central asymmetric units | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | KIF1A / kinesin / motility / microtubule / tubulin / MOTOR PROTEIN | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationneuronal dense core vesicle membrane / dense core granule cytoskeletal transport / anterograde neuronal dense core vesicle transport / retrograde neuronal dense core vesicle transport / regulation of dendritic spine development / cytoskeleton-dependent intracellular transport / regulation of dendritic spine morphogenesis / anterograde axonal transport / Microtubule-dependent trafficking of connexons from Golgi to the plasma membrane / Hedgehog 'off' state ...neuronal dense core vesicle membrane / dense core granule cytoskeletal transport / anterograde neuronal dense core vesicle transport / retrograde neuronal dense core vesicle transport / regulation of dendritic spine development / cytoskeleton-dependent intracellular transport / regulation of dendritic spine morphogenesis / anterograde axonal transport / Microtubule-dependent trafficking of connexons from Golgi to the plasma membrane / Hedgehog 'off' state / Cilium Assembly / Intraflagellar transport / COPI-dependent Golgi-to-ER retrograde traffic / Carboxyterminal post-translational modifications of tubulin / RHOH GTPase cycle / Sealing of the nuclear envelope (NE) by ESCRT-III / Kinesins / PKR-mediated signaling / The role of GTSE1 in G2/M progression after G2 checkpoint / Aggrephagy / Resolution of Sister Chromatid Cohesion / Mitotic Prometaphase / EML4 and NUDC in mitotic spindle formation / Separation of Sister Chromatids / Kinesins / plus-end-directed microtubule motor activity / RHO GTPases activate IQGAPs / RHO GTPases Activate Formins / HSP90 chaperone cycle for steroid hormone receptors (SHR) in the presence of ligand / MHC class II antigen presentation / Recruitment of NuMA to mitotic centrosomes / COPI-mediated anterograde transport / COPI-dependent Golgi-to-ER retrograde traffic / kinesin complex / microtubule-based movement / neuronal dense core vesicle / cytoskeletal motor activity / vesicle-mediated transport / axon cytoplasm / structural constituent of cytoskeleton / microtubule cytoskeleton organization / microtubule cytoskeleton / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / mitotic cell cycle / microtubule binding / microtubule / axon / GTPase activity / dendrite / synapse / GTP binding / perinuclear region of cytoplasm / ATP hydrolysis activity / ATP binding / identical protein binding / metal ion binding / cytoplasm Similarity search - Function | |||||||||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||||||||||||||
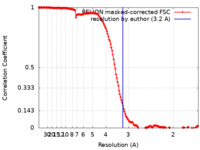
| Method | helical reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||||||||||||||
 Authors Authors | Benoit MPMH / Rao L / Asenjo AB / Gennerich A / Sosa H | |||||||||||||||||||||
| Funding support |  United States, 6 items United States, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Cryo-EM unveils kinesin KIF1A's processivity mechanism and the impact of its pathogenic variant P305L. Authors: Matthieu P M H Benoit / Lu Rao / Ana B Asenjo / Arne Gennerich / Hernando Sosa /  Abstract: Mutations in the microtubule-associated motor protein KIF1A lead to severe neurological conditions known as KIF1A-associated neurological disorders (KAND). Despite insights into its molecular ...Mutations in the microtubule-associated motor protein KIF1A lead to severe neurological conditions known as KIF1A-associated neurological disorders (KAND). Despite insights into its molecular mechanism, high-resolution structures of KIF1A-microtubule complexes remain undefined. Here, we present 2.7-3.5 Å resolution structures of dimeric microtubule-bound KIF1A, including the pathogenic P305L mutant, across various nucleotide states. Our structures reveal that KIF1A binds microtubules in one- and two-heads-bound configurations, with both heads exhibiting distinct conformations with tight inter-head connection. Notably, KIF1A's class-specific loop 12 (K-loop) forms electrostatic interactions with the C-terminal tails of both α- and β-tubulin. The P305L mutation does not disrupt these interactions but alters loop-12's conformation, impairing strong microtubule-binding. Structure-function analysis reveals the K-loop and head-head coordination as major determinants of KIF1A's superprocessive motility. Our findings advance the understanding of KIF1A's molecular mechanism and provide a basis for developing structure-guided therapeutics against KAND. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42544.map.gz emd_42544.map.gz | 38.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42544-v30.xml emd-42544-v30.xml emd-42544.xml emd-42544.xml | 22.7 KB 22.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_42544_fsc.xml emd_42544_fsc.xml | 14.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_42544.png emd_42544.png | 112.4 KB | ||
| Masks |  emd_42544_msk_1.map emd_42544_msk_1.map emd_42544_msk_2.map emd_42544_msk_2.map emd_42544_msk_3.map emd_42544_msk_3.map emd_42544_msk_4.map emd_42544_msk_4.map | 274.6 MB 274.6 MB 274.6 MB 274.6 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-42544.cif.gz emd-42544.cif.gz | 7.5 KB | ||
| Others |  emd_42544_half_map_1.map.gz emd_42544_half_map_1.map.gz emd_42544_half_map_2.map.gz emd_42544_half_map_2.map.gz | 219.1 MB 219.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42544 http://ftp.pdbj.org/pub/emdb/structures/EMD-42544 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42544 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42544 | HTTPS FTP |
-Validation report
| Summary document |  emd_42544_validation.pdf.gz emd_42544_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_42544_full_validation.pdf.gz emd_42544_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_42544_validation.xml.gz emd_42544_validation.xml.gz | 22 KB | Display | |
| Data in CIF |  emd_42544_validation.cif.gz emd_42544_validation.cif.gz | 29.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42544 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42544 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42544 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-42544 | HTTPS FTP |
-Related structure data
| Related structure data |  8utoMC  8utnC  8utpC  8utqC  8utrC  8utsC  8uttC  8utuC  8utvC  8utwC  8utyC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_42544.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42544.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Primary map, locally refined on the two central asymmetric units | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.844 Å | ||||||||||||||||||||||||||||||||||||
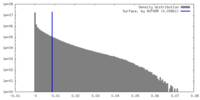
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_42544_msk_1.map emd_42544_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
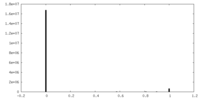
| Density Histograms |
-Mask #2
| File |  emd_42544_msk_2.map emd_42544_msk_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Mask #3
| File |  emd_42544_msk_3.map emd_42544_msk_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
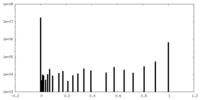
| Density Histograms |
-Mask #4
| File |  emd_42544_msk_4.map emd_42544_msk_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map from the gold Standard refinement
| File | emd_42544_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map from the gold Standard refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
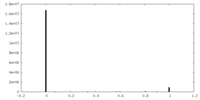
| Density Histograms |
-Half map: Half map from the gold Standard refinement
| File | emd_42544_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map from the gold Standard refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : KIF1A - AMP-PNP two-heads-bound state in complex with a microtubu...
+Supramolecule #1: KIF1A - AMP-PNP two-heads-bound state in complex with a microtubu...
+Supramolecule #2: KIF1A - AMP-PNP
+Supramolecule #3: microtubule
+Macromolecule #1: Kinesin-like protein KIF1A
+Macromolecule #2: Tubulin alpha-1B chain
+Macromolecule #3: Tubulin beta-2B chain
+Macromolecule #4: MAGNESIUM ION
+Macromolecule #5: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER
+Macromolecule #6: GUANOSINE-5'-TRIPHOSPHATE
+Macromolecule #7: GUANOSINE-5'-DIPHOSPHATE
+Macromolecule #8: TAXOL
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: UltrAuFoil / Material: GOLD / Pretreatment - Type: PLASMA CLEANING | ||||||||||||
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.22 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)