+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of Shedu from Bacillus cereus | |||||||||
 Map data Map data | CryoEM structure of Shedu from Bacillus cereus | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Shedu / DUF4263 / Bacterial defense systems / Nuclease / Anti-plasmid defense system / PD-(D/E)XK nuclease / Whirly domain / Two-component signaling / DNA BINDING PROTEIN | |||||||||
| Function / homology | Protein of unknown function DUF4263 / : / Shedu protein SduA, C-terminal / Shedu protein SduA, N-terminal / nuclease activity / defense response to virus / hydrolase activity / Shedu protein SduA Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
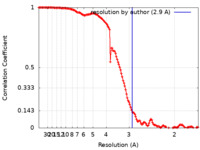
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Gu Y / Corbett K | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2025 Journal: Mol Cell / Year: 2025Title: Bacterial Shedu immune nucleases share a common enzymatic core regulated by diverse sensor domains. Authors: Yajie Gu / Huan Li / Amar Deep / Eray Enustun / Dapeng Zhang / Kevin D Corbett /  Abstract: Prokaryotes possess diverse anti-bacteriophage immune systems, including the single-protein Shedu nuclease. Here, we reveal the structural basis for activation of Bacillus cereus Shedu. Two ...Prokaryotes possess diverse anti-bacteriophage immune systems, including the single-protein Shedu nuclease. Here, we reveal the structural basis for activation of Bacillus cereus Shedu. Two cryoelectron microscopy structures of Shedu show that it switches between inactive and active states through conformational changes affecting active-site architecture, which are controlled by the protein's N-terminal domain (NTD). We find that B. cereus Shedu cleaves near DNA ends with a 3' single-stranded overhang, likely enabling it to specifically degrade the DNA injected by certain bacteriophages. Bioinformatic analysis of Shedu homologs reveals a conserved nuclease domain with remarkably diverse N-terminal regulatory domains: we identify 79 distinct NTD types falling into eight broad classes, including those with predicted nucleic acid binding, enzymatic, and other activities. Together, these data reveal Shedu as a broad family of immune nucleases with a common nuclease core regulated by diverse NTDs that likely respond to a range of signals. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41281.map.gz emd_41281.map.gz | 108.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41281-v30.xml emd-41281-v30.xml emd-41281.xml emd-41281.xml | 19.5 KB 19.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_41281_fsc.xml emd_41281_fsc.xml | 13.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_41281.png emd_41281.png | 113.2 KB | ||
| Filedesc metadata |  emd-41281.cif.gz emd-41281.cif.gz | 6.6 KB | ||
| Others |  emd_41281_half_map_1.map.gz emd_41281_half_map_1.map.gz emd_41281_half_map_2.map.gz emd_41281_half_map_2.map.gz | 200.7 MB 200.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41281 http://ftp.pdbj.org/pub/emdb/structures/EMD-41281 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41281 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41281 | HTTPS FTP |
-Related structure data
| Related structure data |  8ti8MC  8ti9C  8tiaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_41281.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41281.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM structure of Shedu from Bacillus cereus | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.84 Å | ||||||||||||||||||||||||||||||||||||
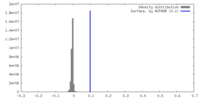
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half Map 1
| File | emd_41281_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
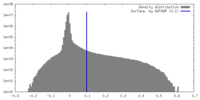
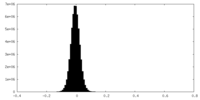
| Density Histograms |
-Half map: Half Map 2
| File | emd_41281_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
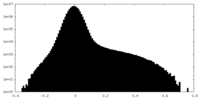
| Density Histograms |
- Sample components
Sample components
-Entire : Bacillus cereus Shedu Tetramer
| Entire | Name: Bacillus cereus Shedu Tetramer |
|---|---|
| Components |
|
-Supramolecule #1: Bacillus cereus Shedu Tetramer
| Supramolecule | Name: Bacillus cereus Shedu Tetramer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 175 KDa |
-Macromolecule #1: Shedu protein SduA
| Macromolecule | Name: Shedu protein SduA / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 44.016969 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SNASDKINVW TTSRDSAVCG DIELKKTSTT RLIFRPEIVN NNKNPKASVR GCFIFQKKGR NALWDDYKEL DMNKLKAEEW IKLEINSDA MLTLTKEIQK HYAVHEKYGV RYGAFHLFKD NPDIEKLIEM FESNTDLLTQ LMEDDKSEAL EKTLEWIVTN D NPDKIIDR ...String: SNASDKINVW TTSRDSAVCG DIELKKTSTT RLIFRPEIVN NNKNPKASVR GCFIFQKKGR NALWDDYKEL DMNKLKAEEW IKLEINSDA MLTLTKEIQK HYAVHEKYGV RYGAFHLFKD NPDIEKLIEM FESNTDLLTQ LMEDDKSEAL EKTLEWIVTN D NPDKIIDR LKNLKEQDLD QLNTLIGIAN LKKVLSVWES NKLTNTSEKF WQSVLKENTW ILSQIFSNPT VLINDEAYVG GK TVKNDSG KLVDFLYANP FSKDAVLIEI KTPSTPLITP TEYRTGVYSA HKDLTGAVTQ VLTYKTTLQR EYQNIDYNNY RQG IKTDFD IITPCCVVIA GMFDTLTDTA HRHSFELYRK ELKNVTVITF DELFERVKGL IKLLEG UniProtKB: Shedu protein SduA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.79 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8.5 Component:
Details: Prepared using deionized water and filtered strelized. | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV | |||||||||||||||
| Details | Freshly collected from size-exclusion column |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Average electron dose: 55.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8ti8: |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)