+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4042 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
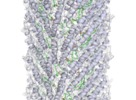
| Title | Cryo-EM structure of the bacterial sex F pilus (pED208) | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | F-pilus Conjugation Type IV secretion system Phospholipid / protein fibril | ||||||||||||
| Function / homology | TraA / TraA / extracellular region / plasma membrane / Pilin Function and homology information Function and homology information | ||||||||||||
| Biological species |   Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria) | ||||||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.6 Å | ||||||||||||
 Authors Authors | Costa TRD / Ilangovan I | ||||||||||||
| Funding support |  United Kingdom, United Kingdom,  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Cell / Year: 2016 Journal: Cell / Year: 2016Title: Structure of the Bacterial Sex F Pilus Reveals an Assembly of a Stoichiometric Protein-Phospholipid Complex. Authors: Tiago R D Costa / Aravindan Ilangovan / Marta Ukleja / Adam Redzej / Joanne M Santini / Terry K Smith / Edward H Egelman / Gabriel Waksman /   Abstract: Conjugative pili are widespread bacterial appendages that play important roles in horizontal gene transfer, in spread of antibiotic resistance genes, and as sites of phage attachment. Among ...Conjugative pili are widespread bacterial appendages that play important roles in horizontal gene transfer, in spread of antibiotic resistance genes, and as sites of phage attachment. Among conjugative pili, the F "sex" pilus encoded by the F plasmid is the best functionally characterized, and it is also historically the most important, as the discovery of F-plasmid-mediated conjugation ushered in the era of molecular biology and genetics. Yet, its structure is unknown. Here, we present atomic models of two F family pili, the F and pED208 pili, generated from cryoelectron microscopy reconstructions at 5.0 and 3.6 Å resolution, respectively. These structures reveal that conjugative pili are assemblies of stoichiometric protein-phospholipid units. We further demonstrate that each pilus type binds preferentially to particular phospholipids. These structures provide the molecular basis for F pilus assembly and also shed light on the remarkable properties of conjugative pili in bacterial secretion and phage infection. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4042.map.gz emd_4042.map.gz | 2.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4042-v30.xml emd-4042-v30.xml emd-4042.xml emd-4042.xml | 10.6 KB 10.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4042.png emd_4042.png | 285.4 KB | ||
| Filedesc metadata |  emd-4042.cif.gz emd-4042.cif.gz | 4.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4042 http://ftp.pdbj.org/pub/emdb/structures/EMD-4042 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4042 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4042 | HTTPS FTP |
-Related structure data
| Related structure data |  5legMC  4044C  4046C  5lerC  5lfbC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4042.map.gz / Format: CCP4 / Size: 16 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4042.map.gz / Format: CCP4 / Size: 16 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : F-pilus (pED208)filament made by an assembly of a stoichiometric ...
| Entire | Name: F-pilus (pED208)filament made by an assembly of a stoichiometric protein-phospholipid complex |
|---|---|
| Components |
|
-Supramolecule #1: F-pilus (pED208)filament made by an assembly of a stoichiometric ...
| Supramolecule | Name: F-pilus (pED208)filament made by an assembly of a stoichiometric protein-phospholipid complex type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 6.3 kDa/nm |
-Macromolecule #1: Pilin
| Macromolecule | Name: Pilin / type: protein_or_peptide / ID: 1 / Number of copies: 80 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria) |
| Molecular weight | Theoretical: 6.75025 KDa |
| Sequence | String: DLLAGGKDDV KATFGADSFV MMCIIIAELI VGVAMYIRTK NLLILLGLVV VIVFTTVGLT FIK UniProtKB: Pilin |
-Macromolecule #2: 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE
| Macromolecule | Name: 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE / type: ligand / ID: 2 / Number of copies: 75 / Formula: LHG |
|---|---|
| Molecular weight | Theoretical: 722.97 Da |
| Chemical component information |  ChemComp-LHG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Lacey / Material: COPPER / Mesh: 400 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.67 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 12.1 Å Applied symmetry - Helical parameters - Δ&Phi: 28.2 ° Applied symmetry - Helical parameters - Axial symmetry: C5 (5 fold cyclic) Resolution.type: BY AUTHOR / Resolution: 3.6 Å / Resolution method: OTHER / Software: (Name: IHRSR (ver. 22), SPIDER) / Number images used: 43952 |
|---|---|
| Startup model | Type of model: NONE |
| Final angle assignment | Type: NOT APPLICABLE / Software - Name: SPIDER (ver. 22) |
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-5leg: |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















