+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tail tube of DT57C bacteriophage in the full state | |||||||||||||||
 Map data Map data | Helical reconstruction of the tail | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Tail tube / T5 / VIRUS / VIRAL PROTEIN | |||||||||||||||
| Function / homology | Bacterial Ig-like domain (group 2) / Invasin/intimin cell-adhesion fragments / Bacterial Ig-like domain 2 / Bacterial Ig-like domain, group 2 / Tail tube protein Function and homology information Function and homology information | |||||||||||||||
| Biological species |  Escherichia phage DT57C (virus) Escherichia phage DT57C (virus) | |||||||||||||||
| Method | helical reconstruction / cryo EM / Resolution: 4.5 Å | |||||||||||||||
 Authors Authors | Ayala R / Moiseenko AV / Chen TH / Kulikov EE / Golomidova AK / Orekhov PS / Street MA / Sokolova OS / Letarov AV / Wolf M | |||||||||||||||
| Funding support |  Russian Federation, Russian Federation,  Japan, 4 items Japan, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Nearly complete structure of bacteriophage DT57C reveals architecture of head-to-tail interface and lateral tail fibers. Authors: Rafael Ayala / Andrey V Moiseenko / Ting-Hua Chen / Eugene E Kulikov / Alla K Golomidova / Philipp S Orekhov / Maya A Street / Olga S Sokolova / Andrey V Letarov / Matthias Wolf /     Abstract: The T5 family of viruses are tailed bacteriophages characterized by a long non-contractile tail. The bacteriophage DT57C is closely related to the paradigmal T5 phage, though it recognizes a ...The T5 family of viruses are tailed bacteriophages characterized by a long non-contractile tail. The bacteriophage DT57C is closely related to the paradigmal T5 phage, though it recognizes a different receptor (BtuB) and features highly divergent lateral tail fibers (LTF). Considerable portions of T5-like phages remain structurally uncharacterized. Here, we present the structure of DT57C determined by cryo-EM, and an atomic model of the virus, which was further explored using all-atom molecular dynamics simulations. The structure revealed a unique way of LTF attachment assisted by a dodecameric collar protein LtfC, and an unusual composition of the phage neck constructed of three protein rings. The tape measure protein (TMP) is organized within the tail tube in a three-stranded parallel α-helical coiled coil which makes direct contact with the genomic DNA. The presence of the C-terminal fragment of the TMP that remains within the tail tip suggests that the tail tip complex returns to its original state after DNA ejection. Our results provide a complete atomic structure of a T5-like phage, provide insights into the process of DNA ejection as well as a structural basis for the design of engineered phages and future mechanistic studies. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34972.map.gz emd_34972.map.gz | 10.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34972-v30.xml emd-34972-v30.xml emd-34972.xml emd-34972.xml | 19.2 KB 19.2 KB | Display Display |  EMDB header EMDB header |
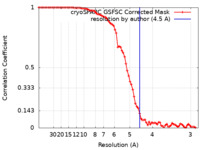
| FSC (resolution estimation) |  emd_34972_fsc.xml emd_34972_fsc.xml | 7.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_34972.png emd_34972.png | 84 KB | ||
| Filedesc metadata |  emd-34972.cif.gz emd-34972.cif.gz | 6.1 KB | ||
| Others |  emd_34972_half_map_1.map.gz emd_34972_half_map_1.map.gz emd_34972_half_map_2.map.gz emd_34972_half_map_2.map.gz | 48 MB 48 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34972 http://ftp.pdbj.org/pub/emdb/structures/EMD-34972 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34972 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34972 | HTTPS FTP |
-Related structure data
| Related structure data |  8hrgMC  8ho3C  8hqkC  8hqoC  8hqzC  8hreC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_34972.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34972.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Helical reconstruction of the tail | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.4 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map A
| File | emd_34972_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_34972_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Escherichia phage DT57C
| Entire | Name:  Escherichia phage DT57C (virus) Escherichia phage DT57C (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Escherichia phage DT57C
| Supramolecule | Name: Escherichia phage DT57C / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 2681606 / Sci species name: Escherichia phage DT57C / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|
-Macromolecule #1: Tail tube protein
| Macromolecule | Name: Tail tube protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Escherichia phage DT57C (virus) Escherichia phage DT57C (virus) |
| Molecular weight | Theoretical: 50.759051 KDa |
| Sequence | String: MSLQLLRNTR IFVSTVKTGH DTTNTQEILV QDDISWGQDS NSTDITVNEA GPRPTRGSKR FNDSLNAAEW SFSTYILPYE DTTTGKQIV PDYMLWHALS SGKAIDLDGD TGAHNNATNF MVNFKDNAYH ELAMLHIYIL TDNAWSYIDS CQINQAEVNV D IEDIGRVT ...String: MSLQLLRNTR IFVSTVKTGH DTTNTQEILV QDDISWGQDS NSTDITVNEA GPRPTRGSKR FNDSLNAAEW SFSTYILPYE DTTTGKQIV PDYMLWHALS SGKAIDLDGD TGAHNNATNF MVNFKDNAYH ELAMLHIYIL TDNAWSYIDS CQINQAEVNV D IEDIGRVT WSGNGNQLIP LDEQPFDPDA LGIDDETYMT IQSSYIKNKL TILKIKDMDS DKEYDIPITG GTFTINNNIT YL TPNIMSR VNIPIGSFTG AFELTGSLTA YLNDKSLGSM ELYKDLVRTL KVVNRFEIAL ILGGEYDDER PAAVLVAKQA HVN IPTIET DDVLGTSVEF KAIPTDLDTG DEGYLGFSNK YTKTTVANLI ATGDGAETPP ILVESITVKS AADATSVTNS DTLQ MSVEV TPPEATNTAV TWSISSGDAA TIDAESGLLT ADVSKTGEVI VKAVAKDGSG VEGTKTITVS AGE UniProtKB: Tail tube protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 67.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)