+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Neck of DT57C bacteriophage in the full state | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Neck / Portal / T5 / VIRUS / VIRAL PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology information: / Head completion protein / : / Bacteriophage Tail Tube Terminator Protein / : / : / Tape measure protein N-terminal / Tape measure protein C-terminal / Bacteriophage/Gene transfer agent portal protein / Phage portal protein Similarity search - Domain/homology | |||||||||||||||
| Biological species |  Escherichia phage DT57C (virus) / Escherichia phage DT57C (virus) /  Tequintavirus DT57C Tequintavirus DT57C | |||||||||||||||
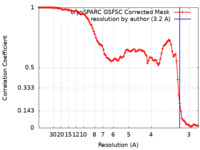
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||||||||
 Authors Authors | Ayala R / Moiseenko AV / Chen TH / Kulikov EE / Golomidova AK / Orekhov PS / Street MA / Sokolova OS / Letarov AV / Wolf M | |||||||||||||||
| Funding support |  Russian Federation, Russian Federation,  Japan, 4 items Japan, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Nearly complete structure of bacteriophage DT57C reveals architecture of head-to-tail interface and lateral tail fibers. Authors: Rafael Ayala / Andrey V Moiseenko / Ting-Hua Chen / Eugene E Kulikov / Alla K Golomidova / Philipp S Orekhov / Maya A Street / Olga S Sokolova / Andrey V Letarov / Matthias Wolf /     Abstract: The T5 family of viruses are tailed bacteriophages characterized by a long non-contractile tail. The bacteriophage DT57C is closely related to the paradigmal T5 phage, though it recognizes a ...The T5 family of viruses are tailed bacteriophages characterized by a long non-contractile tail. The bacteriophage DT57C is closely related to the paradigmal T5 phage, though it recognizes a different receptor (BtuB) and features highly divergent lateral tail fibers (LTF). Considerable portions of T5-like phages remain structurally uncharacterized. Here, we present the structure of DT57C determined by cryo-EM, and an atomic model of the virus, which was further explored using all-atom molecular dynamics simulations. The structure revealed a unique way of LTF attachment assisted by a dodecameric collar protein LtfC, and an unusual composition of the phage neck constructed of three protein rings. The tape measure protein (TMP) is organized within the tail tube in a three-stranded parallel α-helical coiled coil which makes direct contact with the genomic DNA. The presence of the C-terminal fragment of the TMP that remains within the tail tip suggests that the tail tip complex returns to its original state after DNA ejection. Our results provide a complete atomic structure of a T5-like phage, provide insights into the process of DNA ejection as well as a structural basis for the design of engineered phages and future mechanistic studies. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34952.map.gz emd_34952.map.gz | 96.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34952-v30.xml emd-34952-v30.xml emd-34952.xml emd-34952.xml | 19.8 KB 19.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34952_fsc.xml emd_34952_fsc.xml | 11.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_34952.png emd_34952.png | 78.8 KB | ||
| Masks |  emd_34952_msk_1.map emd_34952_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-34952.cif.gz emd-34952.cif.gz | 6.6 KB | ||
| Others |  emd_34952_half_map_1.map.gz emd_34952_half_map_1.map.gz emd_34952_half_map_2.map.gz emd_34952_half_map_2.map.gz | 96.1 MB 96.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34952 http://ftp.pdbj.org/pub/emdb/structures/EMD-34952 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34952 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34952 | HTTPS FTP |
-Related structure data
| Related structure data |  8hqoMC  8ho3C  8hqkC  8hqzC  8hreC  8hrgC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_34952.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34952.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.4 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_34952_msk_1.map emd_34952_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_34952_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_34952_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Tequintavirus DT57C
| Entire | Name:  Tequintavirus DT57C Tequintavirus DT57C |
|---|---|
| Components |
|
-Supramolecule #1: Tequintavirus DT57C
| Supramolecule | Name: Tequintavirus DT57C / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 1567006 / Sci species name: Tequintavirus DT57C / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No |
|---|
-Macromolecule #1: Portal protein
| Macromolecule | Name: Portal protein / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Escherichia phage DT57C (virus) Escherichia phage DT57C (virus) |
| Molecular weight | Theoretical: 45.408574 KDa |
| Sequence | String: MGFKSWITEK LNPGQRIIRD MEPVSHRTNR KPFTTGQAYS KIEILNRTAN MVIDSAAECS YTVGDKYNIV TYANGVKTKT LDTLLNVRP NPFMDISTFR RLVVTDLLFE GCAYIYWDGT SLYHVPAALM QVEADANKFI KKFIFNNQIN YRVDEIIFIK D NSYVCGTN ...String: MGFKSWITEK LNPGQRIIRD MEPVSHRTNR KPFTTGQAYS KIEILNRTAN MVIDSAAECS YTVGDKYNIV TYANGVKTKT LDTLLNVRP NPFMDISTFR RLVVTDLLFE GCAYIYWDGT SLYHVPAALM QVEADANKFI KKFIFNNQIN YRVDEIIFIK D NSYVCGTN SQISGQSRVA TVIDSLEKRS KMLNFKEKFL DNGTVIGLIL ETDEILNKKL RERKQEELQL DYNPSTGQSS VL ILDGGMK AKPYSQISSF KDLDFKEDIA GFNKSICLAF GVPQVLIDGG NNANIRPNIE LFYYMTIIPM LNKLTSSLTF FFG YKITPN TKEVAALTPD KEAEAKHLTS LVNNGIMTGN EARLELNLEP LDDEQMNRIR IPANVAGSAT GVSGQEGGRP QGST EGDKE UniProtKB: Portal protein |
-Macromolecule #2: Head completion protein
| Macromolecule | Name: Head completion protein / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Escherichia phage DT57C (virus) Escherichia phage DT57C (virus) |
| Molecular weight | Theoretical: 19.2039 KDa |
| Sequence | String: MQIITAEDYR LYGGLKRPEL ESGVEMMITA ANALITSLLG MDDADAVDQL INTKPTRKKY FLSSPSATSV TKMTINDKEI DPEQYKLYS DGVILLKFSP PEGYMDVEYT QGGFNPIPED LKLAACMLVD HWHKQDYRQA KTIGGETVTF NNTKSGIPEH I RTIIEVYR RV UniProtKB: Head completion protein |
-Macromolecule #3: Tail terminator protein
| Macromolecule | Name: Tail terminator protein / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Escherichia phage DT57C (virus) Escherichia phage DT57C (virus) |
| Molecular weight | Theoretical: 18.317475 KDa |
| Sequence | String: MDHRTSIAQA LVDRIAQQMD GSQPDEYFNN LYGNVSRQTY KFEEIREFPY VAVHIGTETG QYLPSGQQWM FLELPILVYD KEKTDIQEQ LEKLVADIKT VIDTGGNLEY TVSKPNGSTF PCEATDMSIT SVSTDEGLLA PYGLAEINVT VRYQPPRRSL R R UniProtKB: Tail terminator protein |
-Macromolecule #4: Tape measure protein
| Macromolecule | Name: Tape measure protein / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Escherichia phage DT57C (virus) Escherichia phage DT57C (virus) |
| Molecular weight | Theoretical: 132.626406 KDa |
| Sequence | String: MTDKLIRELL IDVKQKGATR TAKSIENVSD ALENAAAASE LTNEQLGKMP KTLYSIERAA DRAAKSLTKM QASRGMAGIT KSIDSIGAK LDDLSIAMIE VADKLEAGFD GVSRSVKVMG NDVAAATEKV QDRLYDTNRV LGGTARGFND TAGAAGRASR A IGNTSGSA ...String: MTDKLIRELL IDVKQKGATR TAKSIENVSD ALENAAAASE LTNEQLGKMP KTLYSIERAA DRAAKSLTKM QASRGMAGIT KSIDSIGAK LDDLSIAMIE VADKLEAGFD GVSRSVKVMG NDVAAATEKV QDRLYDTNRV LGGTARGFND TAGAAGRASR A IGNTSGSA RGATRDFAAM AKVGGGLPLL YAAIASNIFV LQSAFEQLKL GDQLNRLEEF GVIVGTQTGT PVQSLARSLQ EA AGYAISF EEAMRQASSA SAYGFDAEQL NKFGLVARRA AAVLGVDMTD ALNRVIKGVS KQEIELLDEL GVTIRLNDAY ADY VKQLNA ANTGITYNIN SLTTFQKQQA YANAVIAEST KRFGYLDEVL RATPWEQFAA NADAALRTIQ QAAAKYLGPV IDAI NTVFY TSQASISAEA ARAQEQTNKQ IDPTNVGAVA LSLSASEEGY NKALDMYKES LDKRNKLKSE FDKRMEQADF YTKLA IRQV GEGIPAGLAT AGASEANKKF VEETAAMGLQ VARLDKEVTD STENLNAWKS AYQAAGAAAA KANPEFQKQI NLQRDT TDP GAVYDFNSTV LKGLTEQQKA YNQTKKTASD LANDIQNVAQ NTDTAAKTSA TLADAIKNIE SLSLGTGKSA DEYVKNL NL GYNTLSEMKT ASQALSEYVK LTGNETKNQL AVQQKIADVY NQTKDKEKAQ EAGRRLELQQ LEEQEAALRR VLQTNQGN K AVEREIEKIQ LEKIKLTNQG MEAQKKVKDY TDKILGVDRE IALLNNRTMT DTQYRLAQLN LELTIEKEKY EWYTKQADK QKEAEQSRRA QAQIEREIWK FHQDQQAEMT SKRQEAFENT LTSMFPLAGE MQKMEMQLDF YTQMKELTKD NANEQMRWNA EIAKTRAQM SALTAQRNAQ MQSSVGSSLG AVYTPTTGLS GEDKKFADMG NQLASYDQAI SKLSELNSEA TAVAQSMGNL A NAMIQFSQ GSLDTTSTIA VGMQTVASMI QYSTSQQVSA IDQAIAAEQK RDGKSEASKA KLKKLEAEKL KIQQDAAKKQ II IQTAVAV MQAATAVPYP FSIPLMIAAG LAGALALAQA SSASGMSSIG DSGADTASYL TLGERQKNID VSMSANAGEL SYV RGDKGI GNANSFVPRA EGGNMYPGVS YQMGEHGTEV VTPMVPMKAT PNDELKNSSN STAGRPIILN ISAMDAASFR EFAS SNSGA LRDAVELALN ENGASLKTLG NS UniProtKB: Tape measure protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 67.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)