[English] 日本語
 Yorodumi
Yorodumi- EMDB-33646: Cryo-EM map of IPEC-J2 cell-derived PEDV PT52 S protein with thre... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
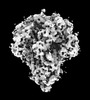
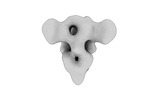
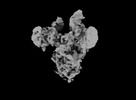
| Title | Cryo-EM map of IPEC-J2 cell-derived PEDV PT52 S protein with three D0-up | |||||||||||||||||||||
 Map data Map data | unsharpened map | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationendocytosis involved in viral entry into host cell / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated virion attachment to host cell / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion membrane / membrane Similarity search - Function | |||||||||||||||||||||
| Biological species |  Porcine epidemic diarrhea virus Porcine epidemic diarrhea virus | |||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||||||||||||||
 Authors Authors | Hsu STD / Draczkowski P / Wang YS | |||||||||||||||||||||
| Funding support |  Taiwan, 6 items Taiwan, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: In situ structure and dynamics of an alphacoronavirus spike protein by cryo-ET and cryo-EM. Authors: Cheng-Yu Huang / Piotr Draczkowski / Yong-Sheng Wang / Chia-Yu Chang / Yu-Chun Chien / Yun-Han Cheng / Yi-Min Wu / Chun-Hsiung Wang / Yuan-Chih Chang / Yen-Chen Chang / Tzu-Jing Yang / Yu-Xi ...Authors: Cheng-Yu Huang / Piotr Draczkowski / Yong-Sheng Wang / Chia-Yu Chang / Yu-Chun Chien / Yun-Han Cheng / Yi-Min Wu / Chun-Hsiung Wang / Yuan-Chih Chang / Yen-Chen Chang / Tzu-Jing Yang / Yu-Xi Tsai / Kay-Hooi Khoo / Hui-Wen Chang / Shang-Te Danny Hsu /   Abstract: Porcine epidemic diarrhea (PED) is a highly contagious swine disease caused by porcine epidemic diarrhea virus (PEDV). PED causes enteric disorders with an exceptionally high fatality in neonates, ...Porcine epidemic diarrhea (PED) is a highly contagious swine disease caused by porcine epidemic diarrhea virus (PEDV). PED causes enteric disorders with an exceptionally high fatality in neonates, bringing substantial economic losses in the pork industry. The trimeric spike (S) glycoprotein of PEDV is responsible for virus-host recognition, membrane fusion, and is the main target for vaccine development and antigenic analysis. The atomic structures of the recombinant PEDV S proteins of two different strains have been reported, but they reveal distinct N-terminal domain 0 (D0) architectures that may correspond to different functional states. The existence of the D0 is a unique feature of alphacoronavirus. Here we combined cryo-electron tomography (cryo-ET) and cryo-electron microscopy (cryo-EM) to demonstrate in situ the asynchronous S protein D0 motions on intact viral particles of a highly virulent PEDV Pintung 52 strain. We further determined the cryo-EM structure of the recombinant S protein derived from a porcine cell line, which revealed additional domain motions likely associated with receptor binding. By integrating mass spectrometry and cryo-EM, we delineated the complex compositions and spatial distribution of the PEDV S protein N-glycans, and demonstrated the functional role of a key N-glycan in modulating the D0 conformation. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33646.map.gz emd_33646.map.gz | 91.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33646-v30.xml emd-33646-v30.xml emd-33646.xml emd-33646.xml | 23 KB 23 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33646_fsc.xml emd_33646_fsc.xml | 12 KB | Display |  FSC data file FSC data file |
| Images |  emd_33646.png emd_33646.png | 34.9 KB | ||
| Others |  emd_33646_additional_1.map.gz emd_33646_additional_1.map.gz emd_33646_half_map_1.map.gz emd_33646_half_map_1.map.gz emd_33646_half_map_2.map.gz emd_33646_half_map_2.map.gz | 173.9 MB 170.8 MB 170.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33646 http://ftp.pdbj.org/pub/emdb/structures/EMD-33646 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33646 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33646 | HTTPS FTP |
-Validation report
| Summary document |  emd_33646_validation.pdf.gz emd_33646_validation.pdf.gz | 831.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_33646_full_validation.pdf.gz emd_33646_full_validation.pdf.gz | 831.4 KB | Display | |
| Data in XML |  emd_33646_validation.xml.gz emd_33646_validation.xml.gz | 20.9 KB | Display | |
| Data in CIF |  emd_33646_validation.cif.gz emd_33646_validation.cif.gz | 27 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33646 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33646 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33646 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33646 | HTTPS FTP |
-Related structure data
| Related structure data |  7y6sMC  7w6mC  7w73C  7y6tC  7y6uC  7y6vC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33646.map.gz / Format: CCP4 / Size: 184 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33646.map.gz / Format: CCP4 / Size: 184 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_33646_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
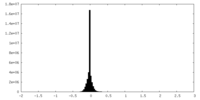
| Density Histograms |
-Half map: #2
| File | emd_33646_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
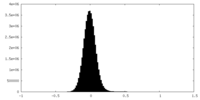
| Density Histograms |
-Half map: #1
| File | emd_33646_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
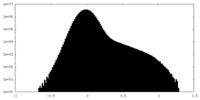
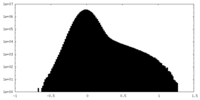
| Density Histograms |
- Sample components
Sample components
-Entire : recombinat Porcine epidemic diarrhea virus (strain Pintung 52) 2P...
| Entire | Name: recombinat Porcine epidemic diarrhea virus (strain Pintung 52) 2P Spike |
|---|---|
| Components |
|
-Supramolecule #1: recombinat Porcine epidemic diarrhea virus (strain Pintung 52) 2P...
| Supramolecule | Name: recombinat Porcine epidemic diarrhea virus (strain Pintung 52) 2P Spike type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: The PEDV PT52 virus was propagated in Vero C1008 cells (ATCC No. CRL-1586) and then inactivated by 2% formaldehyde. |
|---|---|
| Source (natural) | Organism:  Porcine epidemic diarrhea virus / Strain: Pintung 52 Porcine epidemic diarrhea virus / Strain: Pintung 52 |
| Recombinant expression | Organism:  |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Porcine epidemic diarrhea virus Porcine epidemic diarrhea virus |
| Molecular weight | Theoretical: 151.610062 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: LPQDVTRCSA KTNFRRFFSK FNVQAPAVVV LGGYLPIGEN QGVNSTWYCA GQHPTASGVH GIFVSHIRGG HGFEIGISQE PFDPSGYQL YLHKATNGNT NATARLRICQ FPSIKTLGPT ANNDVTIGRN CLFNKAIPAH MSEHSVVGIT WDNDRVTVFS D KIYYFYFK ...String: LPQDVTRCSA KTNFRRFFSK FNVQAPAVVV LGGYLPIGEN QGVNSTWYCA GQHPTASGVH GIFVSHIRGG HGFEIGISQE PFDPSGYQL YLHKATNGNT NATARLRICQ FPSIKTLGPT ANNDVTIGRN CLFNKAIPAH MSEHSVVGIT WDNDRVTVFS D KIYYFYFK NDWSRVATKC YNSGGCAMQY VYEPTYYMLN VTSAGEDGIS YQPCTANCIG YAANVFATEP NGHIPEGFSF NN WFLLSND STLVHGKVVS NQPLLVNCLL AIPKIYGLGQ FFSFNQTIDG VCNGAAVQRA PEALRFNIND TSVILAEGSI VLH TALGTN FSFVCSNSSN PHLATFAIPL GATQVPYYCF FKVDTYNSTV YKFLAVLPPT VREIVITKYG DVYVNGFGYL HLGL LDAVT INFTGHGTDD DVSGFWTIAS TNFVDALIEV QGTAIQRILY CDDPVSQLKC SQVAFDLDDG FYPFSSRNLL SHEQP ISFV TLPSFNAHSF VNITVSASFG GHSGANLIAS DTTINGFSSF CVDTRQFTIS LSYNVTNSYG YVSNSQDSNC PFTLQS VND YLSFSKFCVS TSLLASACTI DLFGYPEFGS GVKFTSLYFQ FTKGELITGT PKPLEGVTDV SFMTLDVCTK YTIYGFK GE GIITLTNSSF LAGVYYTSDS GQLLAFKNVT SGAVYSVTPC SFSEQAAYVD DDIVGVISSL SSSTFNSTRE LPGFFYHS N DGSNCTEPVL VYSNIGVCKS GSIGYVPSQS GQVKIAPTVT GNISIPTNFS MSIRTEYLQL YNTPVSVDCA TYVCNGNSR CKQLLTQYTA ACKTIESALQ LSARLESVEV NSMLTISEEA LQLATISSFN GDGYNFTNVL GVSVYDPARG RVVQKRSFIE DLLFNKVVT NGLGTVDEDY KRCSNGRSVA DLVCAQYYSG VMVLPGVVDA EKLHMYSASL IGGMVLGGFT AAAALPFSYA V QARLNYLA LQTDVLQRNQ QLLAESFNSA IGNITSAFES VKEASSQTSR GLNTVAHALT KVQEVVNSQG AALTQLTVQL QH NFQAISS SIDDIYSRLD PPSADVQVDR LITGRLSALN AFVAQTLTKY TEVQASRKLA QQKVNECVKS QSQRYGFCGG DGE HIFSLV QAAPQGLLFL HTVLVPSDFV DVIAIAGLCV NDEIALTLRE PGLVLFTHEL QNHTATEYFV SSRRMFEPRK PTVS DFVQI ESCVVTYVNL TRDQLPDVIP DYIDVNKTRD EILASLPNRT GPSLPLDVFN ATYLNLTGEI ADLEQRSESL RNTTE ELQS LIYNINNTLV DLEWLNRVET YIKWPEFGSG GYIPEAPRDG QAYVRKDGEW VLLSTFLKGQ DNSADIQHSG RPLESR GPF EQKLISEEDL NMHTGHHHHH H |
-Macromolecule #8: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 8 / Number of copies: 12 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Component:
Details: Blot for 3 seconds before plunging. Force 0. | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV | ||||||||||||
| Details | Cryo-EM map of IPEC-J2 cell-derived PEDV PT52 S protein with three D0-up |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 3701 / Average electron dose: 55.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 64000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-7y6s: |
 Movie
Movie Controller
Controller













 Z
Z Y
Y X
X