+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Human nucleolar pre-60S ribosomal subunit (State A1) - Composite map | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Pre-60S ribosomal subunit / Assembly intermediate / Ribosome / Nucleoprotein complex | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報RNA 2'-O-methyltransferase activity / granular component / RNA metabolic process / positive regulation of protein localization to chromosome, telomeric region / rRNA (uridine-2'-O-ribose)-methyltransferase activity / rRNA (guanine) methyltransferase activity / preribosome binding / lamin filament / regulation of cellular senescence / RNA methylation ...RNA 2'-O-methyltransferase activity / granular component / RNA metabolic process / positive regulation of protein localization to chromosome, telomeric region / rRNA (uridine-2'-O-ribose)-methyltransferase activity / rRNA (guanine) methyltransferase activity / preribosome binding / lamin filament / regulation of cellular senescence / RNA methylation / regulation of megakaryocyte differentiation / regulation of fatty acid biosynthetic process / miRNA-mediated post-transcriptional gene silencing / PeBoW complex / negative regulation of G2/M transition of mitotic cell cycle / positive regulation of protein sumoylation / miRNA-mediated gene silencing by inhibition of translation / rRNA primary transcript binding / maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / blastocyst formation / protein localization to nucleolus / regulation of translation involved in cellular response to UV / negative regulation of formation of translation preinitiation complex / rRNA methylation / ribosomal protein import into nucleus / regulation of G1 to G0 transition / protein-DNA complex disassembly / positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / regulation of glycolytic process / positive regulation of DNA damage response, signal transduction by p53 class mediator / GAIT complex / negative regulation of cell-cell adhesion / TORC2 complex binding / regulation of reactive oxygen species metabolic process / maturation of 5.8S rRNA / stem cell division / G1 to G0 transition / mitotic metaphase chromosome alignment / preribosome, small subunit precursor / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / stem cell population maintenance / rRNA metabolic process / negative regulation of DNA replication / ribosomal large subunit binding / homeostatic process / preribosome, large subunit precursor / macrophage chemotaxis / lung morphogenesis / positive regulation of telomere maintenance / positive regulation of natural killer cell proliferation / rRNA transcription / Protein hydroxylation / Peptide chain elongation / Selenocysteine synthesis / Formation of a pool of free 40S subunits / Eukaryotic Translation Termination / blastocyst development / ubiquitin ligase inhibitor activity / SRP-dependent cotranslational protein targeting to membrane / Response of EIF2AK4 (GCN2) to amino acid deficiency / Viral mRNA Translation / positive regulation of signal transduction by p53 class mediator / ribonucleoprotein complex binding / negative regulation of ubiquitin-dependent protein catabolic process / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / hematopoietic progenitor cell differentiation / GTP hydrolysis and joining of the 60S ribosomal subunit / ribosomal subunit export from nucleus / L13a-mediated translational silencing of Ceruloplasmin expression / Major pathway of rRNA processing in the nucleolus and cytosol / maturation of LSU-rRNA / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / rough endoplasmic reticulum / translation initiation factor activity / MDM2/MDM4 family protein binding / negative regulation of protein ubiquitination / nuclear periphery / cellular response to interleukin-4 / cytosolic ribosome / 転移酵素; 一炭素原子の基を移すもの; メチル基を移すもの / negative regulation of cell migration / regulation of signal transduction by p53 class mediator / cytosolic ribosome assembly / ribosomal large subunit biogenesis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / condensed nuclear chromosome / assembly of large subunit precursor of preribosome / positive regulation of translation / cellular response to estradiol stimulus / mRNA 3'-UTR binding / DNA damage response, signal transduction by p53 class mediator / molecular condensate scaffold activity / cellular response to gamma radiation / response to insulin / bone development / positive regulation of miRNA transcription / cellular response to type II interferon / mRNA 5'-UTR binding / transcription coactivator binding 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.85 Å | |||||||||
 データ登録者 データ登録者 | Vanden Broeck A / Klinge S | |||||||||
| 資金援助 | European Union,  米国, 2件 米国, 2件
| |||||||||
 引用 引用 |  ジャーナル: Science / 年: 2023 ジャーナル: Science / 年: 2023タイトル: Principles of human pre-60 biogenesis. 著者: Arnaud Vanden Broeck / Sebastian Klinge /  要旨: During the early stages of human large ribosomal subunit (60) biogenesis, an ensemble of assembly factors establishes and fine-tunes the essential RNA functional centers of pre-60 particles by an ...During the early stages of human large ribosomal subunit (60) biogenesis, an ensemble of assembly factors establishes and fine-tunes the essential RNA functional centers of pre-60 particles by an unknown mechanism. Here, we report a series of cryo-electron microscopy structures of human nucleolar and nuclear pre-60 assembly intermediates at resolutions of 2.5 to 3.2 angstroms. These structures show how protein interaction hubs tether assembly factor complexes to nucleolar particles and how guanosine triphosphatases and adenosine triphosphatase couple irreversible nucleotide hydrolysis steps to the installation of functional centers. Nuclear stages highlight how a conserved RNA-processing complex, the rixosome, couples large-scale RNA conformational changes with pre-ribosomal RNA processing by the RNA degradation machinery. Our ensemble of human pre-60 particles provides a rich foundation with which to elucidate the molecular principles of ribosome formation. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_29252.map.gz emd_29252.map.gz | 56.4 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-29252-v30.xml emd-29252-v30.xml emd-29252.xml emd-29252.xml | 83 KB 83 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_29252_fsc.xml emd_29252_fsc.xml | 15.8 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_29252.png emd_29252.png | 148.4 KB | ||
| マスクデータ |  emd_29252_msk_1.map emd_29252_msk_1.map | 421.9 MB |  マスクマップ マスクマップ | |
| Filedesc metadata |  emd-29252.cif.gz emd-29252.cif.gz | 19.6 KB | ||
| その他 |  emd_29252_half_map_1.map.gz emd_29252_half_map_1.map.gz emd_29252_half_map_2.map.gz emd_29252_half_map_2.map.gz | 391.1 MB 391.1 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29252 http://ftp.pdbj.org/pub/emdb/structures/EMD-29252 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29252 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29252 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8fkpMC  8fkqC  8fkrC  8fksC  8fktC  8fkuC  8fkvC  8fkwC  8fkxC  8fkyC  8fkzC  8fl0C  8fl2C  8fl3C  8fl4C  8fl6C  8fl7C  8fl9C  8flaC  8flbC  8flcC  8fldC  8fleC  8flfC C: 同じ文献を引用 ( M: このマップから作成された原子モデル |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_29252.map.gz / 形式: CCP4 / 大きさ: 421.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_29252.map.gz / 形式: CCP4 / 大きさ: 421.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.072 Å | ||||||||||||||||||||||||||||||||||||
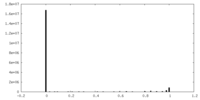
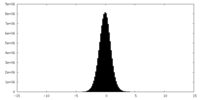
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-マスク #1
| ファイル |  emd_29252_msk_1.map emd_29252_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
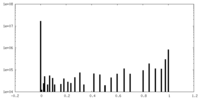
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_29252_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
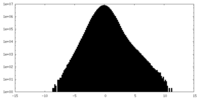
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_29252_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
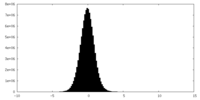
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : Human nucleolar pre-60S ribosomal subunit (State A1)
+超分子 #1: Human nucleolar pre-60S ribosomal subunit (State A1)
+分子 #1: 5.8S rRNA
+分子 #2: ITS2 rRNA
+分子 #3: 28S rRNA
+分子 #4: 60S ribosomal protein L13
+分子 #5: 60S ribosomal protein L13a
+分子 #6: 60S ribosomal protein L14
+分子 #7: 60S ribosomal protein L15
+分子 #8: 60S ribosomal protein L17
+分子 #9: 60S ribosomal protein L18
+分子 #10: 60S ribosomal protein L18a
+分子 #11: 60S ribosomal protein L21
+分子 #12: 60S ribosomal protein L23
+分子 #13: 60S ribosomal protein L23a
+分子 #14: 60S ribosomal protein L26
+分子 #15: 60S ribosomal protein L27a
+分子 #16: 60S ribosomal protein L3
+分子 #17: 60S ribosomal protein L32
+分子 #18: 60S ribosomal protein L35
+分子 #19: 60S ribosomal protein L35a
+分子 #20: 60S ribosomal protein L36
+分子 #21: 60S ribosomal protein L37
+分子 #22: Guanine nucleotide-binding protein-like 3
+分子 #23: Surfeit locus protein 6
+分子 #24: RRP15-like protein
+分子 #25: Protein MAK16 homolog
+分子 #26: Suppressor of SWI4 1 homolog
+分子 #27: Ribosomal RNA processing protein 1 homolog A
+分子 #28: WD repeat-containing protein 74
+分子 #29: Ribosome production factor 1
+分子 #30: 60S ribosomal protein L4
+分子 #31: 60S ribosomal protein L6
+分子 #32: 60S ribosomal protein L7
+分子 #33: 60S ribosomal protein L7a
+分子 #34: MKI67 FHA domain-interacting nucleolar phosphoprotein
+分子 #35: 60S ribosomal protein L7-like 1
+分子 #36: pre-rRNA 2'-O-ribose RNA methyltransferase FTSJ3
+分子 #37: Eukaryotic translation initiation factor 6
+分子 #38: Ribosomal L1 domain-containing protein 1
+分子 #39: Pescadillo homolog
+分子 #40: Probable rRNA-processing protein EBP2
+分子 #41: Ribosome biogenesis protein BRX1 homolog
+分子 #42: GTP-binding protein 4
+分子 #43: Ribosome biogenesis protein BOP1
+分子 #44: Ribosome biogenesis regulatory protein homolog
+分子 #45: Probable ribosome biogenesis protein RLP24
+分子 #46: ATP-dependent RNA helicase DDX18
+分子 #47: Nucleolar protein 16
+分子 #48: MAGNESIUM ION
+分子 #49: ZINC ION
+分子 #50: IRON/SULFUR CLUSTER
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.6 |
|---|---|
| グリッド | モデル: Quantifoil R3.5/1 / 材質: GOLD / メッシュ: 400 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: CONTINUOUS / 支持フィルム - Film thickness: 2 / 前処理 - タイプ: GLOW DISCHARGE / 前処理 - 時間: 30 sec. |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 95 % / チャンバー内温度: 283 K / 装置: FEI VITROBOT MARK IV 詳細: Four applications with manual blotting before last blotting with the Vitrobot.. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 特殊光学系 | エネルギーフィルター - スリット幅: 20 eV |
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 撮影したグリッド数: 4 / 実像数: 172699 / 平均露光時間: 2.0 sec. / 平均電子線量: 60.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.5 µm / 最小 デフォーカス(公称値): 0.5 µm / 倍率(公称値): 64000 |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 精密化 | 空間: REAL |
|---|---|
| 得られたモデル |  PDB-8fkp: |
 ムービー
ムービー コントローラー
コントローラー

















































































































































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)