[English] 日本語
 Yorodumi
Yorodumi- EMDB-28833: Cryo-EM structure of X6 COBRA (H1N1) hemagglutinin bound to CR6261 Fab -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of X6 COBRA (H1N1) hemagglutinin bound to CR6261 Fab | |||||||||
 Map data Map data | Main map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | NIAID hemagglutinin stalk binding antibody / Structural Genomics / Seattle Structural Genomics Center for Infectious Disease / SSGCID / VIRAL PROTEIN / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
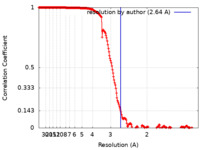
| Method | single particle reconstruction / cryo EM / Resolution: 2.64 Å | |||||||||
 Authors Authors | Seattle Structural Genomics Center for Infectious Disease (SSGCID) | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2024 Journal: Structure / Year: 2024Title: Structural basis for the broad antigenicity of the computationally optimized influenza hemagglutinin X6. Authors: Kaito A Nagashima / John V Dzimianski / Meng Yang / Jan Abendroth / Giuseppe A Sautto / Ted M Ross / Rebecca M DuBois / Thomas E Edwards / Jarrod J Mousa /  Abstract: Influenza causes significant morbidity and mortality. As an alternative approach to current seasonal vaccines, the computationally optimized broadly reactive antigen (COBRA) platform has been ...Influenza causes significant morbidity and mortality. As an alternative approach to current seasonal vaccines, the computationally optimized broadly reactive antigen (COBRA) platform has been previously applied to hemagglutinin (HA). This approach integrates wild-type HA sequences into a single immunogen to expand the breadth of accessible antibody epitopes. Adding to previous studies of H1, H3, and H5 COBRA HAs, we define the structural features of another H1 subtype COBRA, X6, that incorporates HA sequences from before and after the 2009 H1N1 influenza pandemic. We determined structures of this antigen alone and in complex with COBRA-specific as well as broadly reactive and functional antibodies, analyzing its antigenicity. We found that X6 possesses features representing both historic and recent H1 HA strains, enabling binding to both head- and stem-reactive antibodies. Overall, these data confirm the integrity of broadly reactive antibody epitopes of X6 and contribute to design efforts for a next-generation vaccine. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28833.map.gz emd_28833.map.gz | 340.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28833-v30.xml emd-28833-v30.xml emd-28833.xml emd-28833.xml | 20.2 KB 20.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28833_fsc.xml emd_28833_fsc.xml | 19.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_28833.png emd_28833.png | 132 KB | ||
| Filedesc metadata |  emd-28833.cif.gz emd-28833.cif.gz | 6.8 KB | ||
| Others |  emd_28833_half_map_1.map.gz emd_28833_half_map_1.map.gz emd_28833_half_map_2.map.gz emd_28833_half_map_2.map.gz | 614.9 MB 614.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28833 http://ftp.pdbj.org/pub/emdb/structures/EMD-28833 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28833 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28833 | HTTPS FTP |
-Related structure data
| Related structure data |  8f38MC  8ghkC  8sj9C  8v7oC M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28833.map.gz / Format: CCP4 / Size: 662.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28833.map.gz / Format: CCP4 / Size: 662.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Main map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.675 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: handedness corrected half map B
| File | emd_28833_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | handedness corrected half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: handedness corrected half map A
| File | emd_28833_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | handedness corrected half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of X6 COBRA (H1N1) hemagglutinin bound to CR6261 Fab
| Entire | Name: Cryo-EM structure of X6 COBRA (H1N1) hemagglutinin bound to CR6261 Fab |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of X6 COBRA (H1N1) hemagglutinin bound to CR6261 Fab
| Supramolecule | Name: Cryo-EM structure of X6 COBRA (H1N1) hemagglutinin bound to CR6261 Fab type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Molecular weight | Theoretical: 450 KDa |
-Macromolecule #1: CR6261 Fab light chain
| Macromolecule | Name: CR6261 Fab light chain / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.257783 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSVLTQPPSV SAAPGQKVTI SCSGSSSNIG NDYVSWYQQL PGTAPKLLIY DNNKRPSGIP DRFSGSKSGT SATLGITGLQ TGDEANYYC ATWDRRPTAY VVFGGGTKLT VLGAAAGQPK AAPSVTLFPP SSEELQANKA TLVCLISDFY PGAVTVAWKA D SSPVKAGV ...String: QSVLTQPPSV SAAPGQKVTI SCSGSSSNIG NDYVSWYQQL PGTAPKLLIY DNNKRPSGIP DRFSGSKSGT SATLGITGLQ TGDEANYYC ATWDRRPTAY VVFGGGTKLT VLGAAAGQPK AAPSVTLFPP SSEELQANKA TLVCLISDFY PGAVTVAWKA D SSPVKAGV ETTTPSKQSN NKYAASSYLS LTPEQWKSHR SYSCQVTHEG STVEKTVAPT ECS |
-Macromolecule #2: CR6261 Fab heavy chain
| Macromolecule | Name: CR6261 Fab heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.810018 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLVESGAE VKKPGSSVKV SCKASGGPFR SYAISWVRQA PGQGPEWMGG IIPIFGTTKY APKFQGRVTI TADDFAGTVY MELSSLRSE DTAMYYCAKH MGYQVRETMD VWGKGTTVTV SSASTKGPSV FPLAPSSKST SGGTAALGCL VKDYFPEPVT V SWNSGALT ...String: EVQLVESGAE VKKPGSSVKV SCKASGGPFR SYAISWVRQA PGQGPEWMGG IIPIFGTTKY APKFQGRVTI TADDFAGTVY MELSSLRSE DTAMYYCAKH MGYQVRETMD VWGKGTTVTV SSASTKGPSV FPLAPSSKST SGGTAALGCL VKDYFPEPVT V SWNSGALT SGVHTFPAVL QSSGLYSLSS VVTVPSSSLG TQTYICNVNH KPSNTKVDKR VEPKSCDKHH HHHH |
-Macromolecule #3: hemagglutinin
| Macromolecule | Name: hemagglutinin / type: protein_or_peptide / ID: 3 / Details: X6 COBRA (H1N1) hemagglutinin construct / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 63.171098 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MEARLLVLLC AFAATNADTI CIGYHANNST DTVDTVLEKN VTVTHSVNLL EDSHNGKLCL LKGIAPLQLG NCSVAGWILG NPECELLIS KESWSYIVET PNPENGTCYP GYFADYEELR EQLSSVSSFE RFEIFPKESS WPNHTVTGVS ASCSHNGKSS F YRNLLWLT ...String: MEARLLVLLC AFAATNADTI CIGYHANNST DTVDTVLEKN VTVTHSVNLL EDSHNGKLCL LKGIAPLQLG NCSVAGWILG NPECELLIS KESWSYIVET PNPENGTCYP GYFADYEELR EQLSSVSSFE RFEIFPKESS WPNHTVTGVS ASCSHNGKSS F YRNLLWLT GKNGLYPNLS KSYANNKEKE VLVLWGVHHP PNIGDQRALY HTENAYVSVV SSHYSRKFTP EIAKRPKVRD QE GRINYYW TLLEPGDTII FEANGNLIAP RYAFALSRGF GSGIITSNAP MDECDAKCQT PQGAINSSLP FQNVHPVTIG ECP KYVRSA KLRMVTGLRN IPSIQSRGLF GAIAGFIEGG WTGMVDGWYG YHHQNEQGSG YAADQKSTQN AINGITNKVN SVIE KMNTQ FTAVGKEFNK LERRMENLNK KVDDGFLDIW TYNAELLVLL ENERTLDFHD SNVKNLYEKV KSQLKNNAKE IGNGC FEFY HKCNNECMES VKNGTYDYPK YSEESKLNRE KIDGVKLESM GVYQILAIYS TVASSLVLLV SLGAISFWMC SNGSLQ CRI CI |
-Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 6 / Number of copies: 18 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #7: water
| Macromolecule | Name: water / type: ligand / ID: 7 / Number of copies: 50 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Model: EMS Lacey Carbon / Material: COPPER / Mesh: 300 / Support film - Material: GRAPHENE OXIDE / Support film - topology: LACEY | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 8973 / Average exposure time: 2.4 sec. / Average electron dose: 80.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)