登録情報 データベース : EMDB / ID : EMD-28487タイトル Eag Kv channel with voltage sensor in the up conformation 複合体 : Complex of Eag Kv channel bound to the inhibitor calmodulin-Ca2+タンパク質・ペプチド : Potassium voltage-gated channel subfamily H member 1タンパク質・ペプチド : Calmodulin-1 / 機能・相同性 分子機能 ドメイン・相同性 構成要素
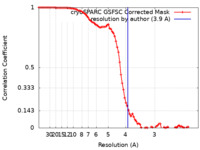
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Rattus norvegicus (ドブネズミ) / Homo sapiens (ヒト)手法 / / 解像度 : 3.9 Å Mandala VS / MacKinnon R 資金援助 Organization Grant number 国 Howard Hughes Medical Institute (HHMI)
ジャーナル : Proc Natl Acad Sci U S A / 年 : 2022タイトル : Voltage-sensor movements in the Eag Kv channel under an applied electric field.著者 : Venkata Shiva Mandala / Roderick MacKinnon / 要旨 : Voltage-dependent ion channels regulate the opening of their pores by sensing the membrane voltage. This process underlies the propagation of action potentials and other forms of electrical activity ... Voltage-dependent ion channels regulate the opening of their pores by sensing the membrane voltage. This process underlies the propagation of action potentials and other forms of electrical activity in cells. The voltage dependence of these channels is governed by the transmembrane displacement of the positive charged S4 helix within their voltage-sensor domains. We use cryo-electron microscopy to visualize this movement in the mammalian Eag voltage-dependent potassium channel in lipid membrane vesicles with a voltage difference across the membrane. Multiple structural configurations show that the applied electric field displaces S4 toward the cytoplasm by two helical turns, resulting in an extended interfacial helix near the inner membrane leaflet. The position of S4 in this down conformation is sterically incompatible with an open pore, thus explaining how movement of the voltage sensor at hyperpolarizing membrane voltages locks the pore shut in this kind of voltage-dependent K (K) channel. The structures solved in lipid bilayer vesicles detail the intricate interplay between K channels and membranes, from showing how arginines are stabilized deep within the membrane and near phospholipid headgroups, to demonstrating how the channel reshapes the inner leaflet of the membrane itself. 履歴 登録 2022年10月4日 - ヘッダ(付随情報) 公開 2022年11月16日 - マップ公開 2022年11月16日 - 更新 2025年6月4日 - 現状 2025年6月4日 処理サイト : RCSB / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
基本情報
 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報
 Homo sapiens (ヒト)
Homo sapiens (ヒト) データ登録者
データ登録者 米国, 1件
米国, 1件  引用
引用 ジャーナル: Proc Natl Acad Sci U S A / 年: 2022
ジャーナル: Proc Natl Acad Sci U S A / 年: 2022
 構造の表示
構造の表示 ダウンロードとリンク
ダウンロードとリンク emd_28487.map.gz
emd_28487.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-28487-v30.xml
emd-28487-v30.xml emd-28487.xml
emd-28487.xml EMDBヘッダ
EMDBヘッダ emd_28487_fsc.xml
emd_28487_fsc.xml FSCデータファイル
FSCデータファイル emd_28487.png
emd_28487.png emd-28487.cif.gz
emd-28487.cif.gz emd_28487_additional_1.map.gz
emd_28487_additional_1.map.gz emd_28487_half_map_1.map.gz
emd_28487_half_map_1.map.gz emd_28487_half_map_2.map.gz
emd_28487_half_map_2.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-28487
http://ftp.pdbj.org/pub/emdb/structures/EMD-28487 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28487
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28487 emd_28487_validation.pdf.gz
emd_28487_validation.pdf.gz EMDB検証レポート
EMDB検証レポート emd_28487_full_validation.pdf.gz
emd_28487_full_validation.pdf.gz emd_28487_validation.xml.gz
emd_28487_validation.xml.gz emd_28487_validation.cif.gz
emd_28487_validation.cif.gz https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28487
https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28487 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28487
ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28487 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_28487.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_28487.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素

 Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 ムービー
ムービー コントローラー
コントローラー




































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)