[English] 日本語
 Yorodumi
Yorodumi- EMDB-27803: Complex between MLL1-WRAD and an H2B-ubiquitinated nucleosome- State 3 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Complex between MLL1-WRAD and an H2B-ubiquitinated nucleosome- State 3 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) / | |||||||||
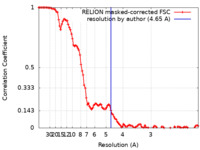
| Method | single particle reconstruction / cryo EM / Resolution: 4.65 Å | |||||||||
 Authors Authors | Niklas HA / Rahman S / Worden EJ / Wolberger C | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: Multistate structures of the MLL1-WRAD complex bound to H2B-ubiquitinated nucleosome. Authors: Sanim Rahman / Niklas A Hoffmann / Evan J Worden / Marissa L Smith / Kevin E W Namitz / Bruce A Knutson / Michael S Cosgrove / Cynthia Wolberger /  Abstract: The human Mixed Lineage Leukemia-1 (MLL1) complex methylates histone H3K4 to promote transcription and is stimulated by monoubiquitination of histone H2B. Recent structures of the MLL1-WRAD core ...The human Mixed Lineage Leukemia-1 (MLL1) complex methylates histone H3K4 to promote transcription and is stimulated by monoubiquitination of histone H2B. Recent structures of the MLL1-WRAD core complex, which comprises the MLL1 methyltransferase, DR5, bBp5, sh2L, and PY-30, have revealed variability in the docking of MLL1-WRAD on nucleosomes. In addition, portions of the Ash2L structure and the position of DPY30 remain ambiguous. We used an integrated approach combining cryoelectron microscopy (cryo-EM) and mass spectrometry cross-linking to determine a structure of the MLL1-WRAD complex bound to ubiquitinated nucleosomes. The resulting model contains the Ash2L intrinsically disordered region (IDR), SPRY insertion region, Sdc1-DPY30 interacting region (SDI-motif), and the DPY30 dimer. We also resolved three additional states of MLL1-WRAD lacking one or more subunits, which may reflect different steps in the assembly of MLL1-WRAD. The docking of subunits in all four states differs from structures of MLL1-WRAD bound to unmodified nucleosomes, suggesting that H2B-ubiquitin favors assembly of the active complex. Our results provide a more complete picture of MLL1-WRAD and the role of ubiquitin in promoting formation of the active methyltransferase complex. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27803.map.gz emd_27803.map.gz | 8.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27803-v30.xml emd-27803-v30.xml emd-27803.xml emd-27803.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27803_fsc.xml emd_27803_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_27803.png emd_27803.png | 51.6 KB | ||
| Masks |  emd_27803_msk_1.map emd_27803_msk_1.map | 103 MB |  Mask map Mask map | |
| Others |  emd_27803_additional_1.map.gz emd_27803_additional_1.map.gz emd_27803_half_map_1.map.gz emd_27803_half_map_1.map.gz emd_27803_half_map_2.map.gz emd_27803_half_map_2.map.gz | 80.5 MB 80.8 MB 80.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27803 http://ftp.pdbj.org/pub/emdb/structures/EMD-27803 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27803 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27803 | HTTPS FTP |
-Validation report
| Summary document |  emd_27803_validation.pdf.gz emd_27803_validation.pdf.gz | 682 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27803_full_validation.pdf.gz emd_27803_full_validation.pdf.gz | 681.6 KB | Display | |
| Data in XML |  emd_27803_validation.xml.gz emd_27803_validation.xml.gz | 17.9 KB | Display | |
| Data in CIF |  emd_27803_validation.cif.gz emd_27803_validation.cif.gz | 23.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27803 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27803 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27803 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27803 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27803.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27803.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.058 Å | ||||||||||||||||||||||||||||||||||||
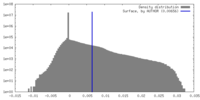
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27803_msk_1.map emd_27803_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unsharpened map
| File | emd_27803_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_27803_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_27803_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : MLL1 WRAD complex bound to the ubiquitinated nucleosome
| Entire | Name: MLL1 WRAD complex bound to the ubiquitinated nucleosome |
|---|---|
| Components |
|
-Supramolecule #1: MLL1 WRAD complex bound to the ubiquitinated nucleosome
| Supramolecule | Name: MLL1 WRAD complex bound to the ubiquitinated nucleosome type: complex / Chimera: Yes / ID: 1 / Parent: 0 |
|---|
-Supramolecule #2: MLL1 WRAD complex
| Supramolecule | Name: MLL1 WRAD complex / type: complex / Chimera: Yes / ID: 2 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
-Supramolecule #3: nucleosome
| Supramolecule | Name: nucleosome / type: complex / Chimera: Yes / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism: |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 5091 / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.8000000000000003 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)