[English] 日本語
 Yorodumi
Yorodumi- EMDB-27699: Cryo-EM structure of Arabidopsis SPY in complex with GDP-fucose -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Arabidopsis SPY in complex with GDP-fucose | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | O-fucosyltransferase / SPY / TRANSFERASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationprotein N-acetylglucosaminyltransferase activity / negative regulation of gibberellic acid mediated signaling pathway / peptide-O-fucosyltransferase / protein O-linked glycosylation via fucose / peptide-O-fucosyltransferase activity / gibberellic acid mediated signaling pathway / cytokinin-activated signaling pathway / protein O-GlcNAc transferase / protein O-acetylglucosaminyltransferase activity / flower development ...protein N-acetylglucosaminyltransferase activity / negative regulation of gibberellic acid mediated signaling pathway / peptide-O-fucosyltransferase / protein O-linked glycosylation via fucose / peptide-O-fucosyltransferase activity / gibberellic acid mediated signaling pathway / cytokinin-activated signaling pathway / protein O-GlcNAc transferase / protein O-acetylglucosaminyltransferase activity / flower development / regulation of reactive oxygen species metabolic process / rhythmic process / cell differentiation / nucleus / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
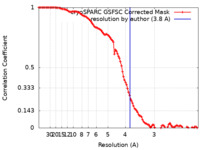
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Kumar S / Zhou Y / Dillard L / Borgnia MJ / Bartesaghi A / Zhou P | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Cryo-EM structure of the full length Arabidopsis SPY with complete TPRs Authors: Kumar S / Zhou Y | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27699.map.gz emd_27699.map.gz | 117 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27699-v30.xml emd-27699-v30.xml emd-27699.xml emd-27699.xml | 20.6 KB 20.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27699_fsc.xml emd_27699_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_27699.png emd_27699.png | 79 KB | ||
| Filedesc metadata |  emd-27699.cif.gz emd-27699.cif.gz | 6.5 KB | ||
| Others |  emd_27699_additional_1.map.gz emd_27699_additional_1.map.gz emd_27699_additional_2.map.gz emd_27699_additional_2.map.gz emd_27699_additional_3.map.gz emd_27699_additional_3.map.gz | 117.9 MB 118.2 MB 118 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27699 http://ftp.pdbj.org/pub/emdb/structures/EMD-27699 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27699 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27699 | HTTPS FTP |
-Related structure data
| Related structure data |  8dtiMC  8dtfC  8dtgC  8dthC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27699.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27699.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Overall lower resolution map
| File | emd_27699_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Overall lower resolution map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Focused component map (monomer B)
| File | emd_27699_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Focused component map (monomer B) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Focused component map (monomer A)
| File | emd_27699_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Focused component map (monomer A) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SPY in complex with GDP-fucose
| Entire | Name: SPY in complex with GDP-fucose |
|---|---|
| Components |
|
-Supramolecule #1: SPY in complex with GDP-fucose
| Supramolecule | Name: SPY in complex with GDP-fucose / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Probable UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltra...
| Macromolecule | Name: Probable UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase SPINDLY type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: protein O-GlcNAc transferase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 105.278859 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MVGLEDDTER ERSPVVENGF SNGSRSSSSS AGVLSPSRKV TQGNDTLSYA NILRARNKFA DALALYEAML EKDSKNVEAH IGKGICLQT QNKGNLAFDC FSEAIRLDPH NACALTHCGI LHKEEGRLVE AAESYQKALM ADASYKPAAE CLAIVLTDLG T SLKLAGNT ...String: MVGLEDDTER ERSPVVENGF SNGSRSSSSS AGVLSPSRKV TQGNDTLSYA NILRARNKFA DALALYEAML EKDSKNVEAH IGKGICLQT QNKGNLAFDC FSEAIRLDPH NACALTHCGI LHKEEGRLVE AAESYQKALM ADASYKPAAE CLAIVLTDLG T SLKLAGNT QEGIQKYYEA LKIDPHYAPA YYNLGVVYSE MMQYDNALSC YEKAALERPM YAEAYCNMGV IYKNRGDLEM AI TCYERCL AVSPNFEIAK NNMAIALTDL GTKVKLEGDV TQGVAYYKKA LYYNWHYADA MYNLGVAYGE MLKFDMAIVF YEL AFHFNP HCAEACNNLG VLYKDRDNLD KAVECYQMAL SIKPNFAQSL NNLGVVYTVQ GKMDAAASMI EKAILANPTY AEAF NNLGV LYRDAGNITM AIDAYEECLK IDPDSRNAGQ NRLLAMNYIN EGLDDKLFEA HRDWGWRFTR LHPQYTSWDN LKDPE RPIT IGYISPDFFT HSVSYFIEAP LTHHDYTKYK VVVYSAVVKA DAKTYRFRDK VLKKGGVWKD IYGIDEKKIA SMVRED KID ILVELTGHTA NNKLGTMACR PAPVQVTWIG YPNTTGLPTV DYRITDSLAD PPDTKQKQVE ELVRLPDCFL CYTPSPE AG PVCPTPALSN GFVTFGSFNN LAKITPKVLQ VWARILCAVP NSRLVVKCKP FCCDSIRQRF LTTLEQLGLE SKRVDLLP L ILFNHDHMQA YSLMDISLDT FPYAGTTTTC ESLYMGVPCV TMAGSVHAHN VGVSLLTKVG LGHLVAKNED EYVQLSVDL ASDVTALSKL RMSLRDLMAG SPVCNGPSFA VGLESAYRNM WKKYCKGEVP SLRRMEMLQK EVHDDPLISK DLGPSRVSVT GEATPSLKA NGSAPVPSSL PTQSPQLSKR MDSTSGGSEN LYFQGGSHHH HHHHHHHGGW SHPQFEK UniProtKB: Probable UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase SPINDLY |
-Macromolecule #2: GUANOSINE-5'-DIPHOSPHATE-BETA-L-FUCOPYRANOSE
| Macromolecule | Name: GUANOSINE-5'-DIPHOSPHATE-BETA-L-FUCOPYRANOSE / type: ligand / ID: 2 / Number of copies: 2 / Formula: GFB |
|---|---|
| Molecular weight | Theoretical: 589.342 Da |
| Chemical component information |  ChemComp-GFB: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: C-flat-1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: PLASMA CLEANING |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 52.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)