[English] 日本語
 Yorodumi
Yorodumi- EMDB-26949: Local refinement of RhAG/CE trimer, class 1 of erythrocyte ankyri... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Local refinement of RhAG/CE trimer, class 1 of erythrocyte ankyrin-1 complex | |||||||||
 Map data Map data | Main map used for model building/refinement. Density modified and cropped using phenix.resolve_cryo_em, resampled on fine grid using relion_image_handler. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Membrane Protein / ankyrin complex / Erythrocyte / STRUCTURAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmethylammonium transmembrane transport / Defective RHAG causes regulator type Rh-null hemolytic anemia (RHN) / Rhesus blood group biosynthesis / methylammonium transmembrane transporter activity / Rhesus glycoproteins mediate ammonium transport / carbon dioxide transmembrane transport / carbon dioxide transmembrane transporter activity / ammonium homeostasis / spectrin-associated cytoskeleton / maintenance of epithelial cell apical/basal polarity ...methylammonium transmembrane transport / Defective RHAG causes regulator type Rh-null hemolytic anemia (RHN) / Rhesus blood group biosynthesis / methylammonium transmembrane transporter activity / Rhesus glycoproteins mediate ammonium transport / carbon dioxide transmembrane transport / carbon dioxide transmembrane transporter activity / ammonium homeostasis / spectrin-associated cytoskeleton / maintenance of epithelial cell apical/basal polarity / NrCAM interactions / Neurofascin interactions / ammonium transmembrane transport / leak channel activity / ammonium channel activity / ankyrin-1 complex / intracellular monoatomic ion homeostasis / CHL1 interactions / cytoskeletal anchor activity / bicarbonate transport / M band / : / Interaction between L1 and Ankyrins / ankyrin binding / spectrin binding / exocytosis / axolemma / endoplasmic reticulum to Golgi vesicle-mediated transport / erythrocyte development / COPI-mediated anterograde transport / cytoskeleton organization / sarcoplasmic reticulum / protein localization to plasma membrane / carbon dioxide transport / Erythrocytes take up oxygen and release carbon dioxide / Erythrocytes take up carbon dioxide and release oxygen / sarcolemma / multicellular organismal-level iron ion homeostasis / structural constituent of cytoskeleton / cytoplasmic side of plasma membrane / Z disc / ATPase binding / protein phosphatase binding / basolateral plasma membrane / cytoskeleton / transmembrane transporter binding / postsynaptic membrane / neuron projection / structural molecule activity / enzyme binding / signal transduction / membrane / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.5 Å | |||||||||
 Authors Authors | Vallese F / Kim K / Yen LY / Johnston JD / Noble AJ / Cali T / Clarke OB | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2022 Journal: Nat Struct Mol Biol / Year: 2022Title: Architecture of the human erythrocyte ankyrin-1 complex. Authors: Francesca Vallese / Kookjoo Kim / Laura Y Yen / Jake D Johnston / Alex J Noble / Tito Calì / Oliver Biggs Clarke /   Abstract: The stability and shape of the erythrocyte membrane is provided by the ankyrin-1 complex, but how it tethers the spectrin-actin cytoskeleton to the lipid bilayer and the nature of its association ...The stability and shape of the erythrocyte membrane is provided by the ankyrin-1 complex, but how it tethers the spectrin-actin cytoskeleton to the lipid bilayer and the nature of its association with the band 3 anion exchanger and the Rhesus glycoproteins remains unknown. Here we present structures of ankyrin-1 complexes purified from human erythrocytes. We reveal the architecture of a core complex of ankyrin-1, the Rhesus proteins RhAG and RhCE, the band 3 anion exchanger, protein 4.2, glycophorin A and glycophorin B. The distinct T-shaped conformation of membrane-bound ankyrin-1 facilitates recognition of RhCE and, unexpectedly, the water channel aquaporin-1. Together, our results uncover the molecular details of ankyrin-1 association with the erythrocyte membrane, and illustrate the mechanism of ankyrin-mediated membrane protein clustering. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26949.map.gz emd_26949.map.gz | 76.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26949-v30.xml emd-26949-v30.xml emd-26949.xml emd-26949.xml | 36.3 KB 36.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26949_fsc.xml emd_26949_fsc.xml | 15.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_26949.png emd_26949.png | 68.9 KB | ||
| Filedesc metadata |  emd-26949.cif.gz emd-26949.cif.gz | 8.1 KB | ||
| Others |  emd_26949_additional_1.map.gz emd_26949_additional_1.map.gz emd_26949_additional_2.map.gz emd_26949_additional_2.map.gz emd_26949_additional_3.map.gz emd_26949_additional_3.map.gz emd_26949_additional_4.map.gz emd_26949_additional_4.map.gz emd_26949_half_map_1.map.gz emd_26949_half_map_1.map.gz emd_26949_half_map_2.map.gz emd_26949_half_map_2.map.gz | 328 MB 322.7 MB 322.6 MB 617.3 KB 75.3 MB 75.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26949 http://ftp.pdbj.org/pub/emdb/structures/EMD-26949 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26949 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26949 | HTTPS FTP |
-Related structure data
| Related structure data |  7v0sMC  7uz3C  7uzeC  7uzqC  7uzsC  7uzuC  7uzvC  7v07C  7v0kC  7v0mC  7v0qC  7v0tC  7v0uC  7v0xC  7v0yC  7v19C  8crqC  8crrC  8crtC  8cs9C  8cslC  8csvC  8cswC  8csxC  8csyC  8ct2C  8ct3C  8cteC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26949.map.gz / Format: CCP4 / Size: 82 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26949.map.gz / Format: CCP4 / Size: 82 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Main map used for model building/refinement. Density modified and cropped using phenix.resolve_cryo_em, resampled on fine grid using relion_image_handler. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.415 Å | ||||||||||||||||||||||||||||||||||||
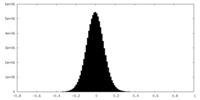
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Sharpened map (B=58).
| File | emd_26949_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map (B=58). | ||||||||||||
| Projections & Slices |
| ||||||||||||
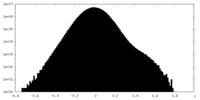
| Density Histograms |
-Additional map: Half map 1 (unmodified).
| File | emd_26949_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 (unmodified). | ||||||||||||
| Projections & Slices |
| ||||||||||||
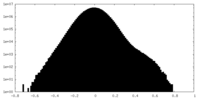
| Density Histograms |
-Additional map: Half map 2 (unmodified).
| File | emd_26949_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 (unmodified). | ||||||||||||
| Projections & Slices |
| ||||||||||||
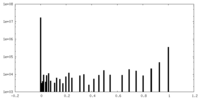
| Density Histograms |
-Additional map: Mask used for FSC calculation.
| File | emd_26949_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mask used for FSC calculation. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1 (cropped and resampled).
| File | emd_26949_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 (cropped and resampled). | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2 (cropped and resampled).
| File | emd_26949_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 (cropped and resampled). | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Erythrocyte ankyrin-1 complex
| Entire | Name: Erythrocyte ankyrin-1 complex |
|---|---|
| Components |
|
-Supramolecule #1: Erythrocyte ankyrin-1 complex
| Supramolecule | Name: Erythrocyte ankyrin-1 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 Details: Purified by density gradient centrifugation and size exclusion chromatography from digitonin-solubilized erythrocyte membranes. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organ: Blood / Tissue: Erythrocytes / Location in cell: Plasma membrane Homo sapiens (human) / Organ: Blood / Tissue: Erythrocytes / Location in cell: Plasma membrane |
-Macromolecule #1: Blood group Rh(CE) polypeptide
| Macromolecule | Name: Blood group Rh(CE) polypeptide / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 45.598918 KDa |
| Sequence | String: MSSKYPRSVR RCLPLWALTL EAALILLFYF FTHYDASLED QKGLVASYQV GQDLTVMAAL GLGFLTSNFR RHSWSSVAFN LFMLALGVQ WAILLDGFLS QFPPGKVVIT LFSIRLATMS AMSVLISAGA VLGKVNLAQL VVMVLVEVTA LGTLRMVISN I FNTDYHMN ...String: MSSKYPRSVR RCLPLWALTL EAALILLFYF FTHYDASLED QKGLVASYQV GQDLTVMAAL GLGFLTSNFR RHSWSSVAFN LFMLALGVQ WAILLDGFLS QFPPGKVVIT LFSIRLATMS AMSVLISAGA VLGKVNLAQL VVMVLVEVTA LGTLRMVISN I FNTDYHMN LRHFYVFAAY FGLTVAWCLP KPLPKGTEDN DQRATIPSLS AMLGALFLWM FWPSVNSPLL RSPIQRKNAM FN TYYALAV SVVTAISGSS LAHPQRKISM TYVHSAVLAG GVAVGTSCHL IPSPWLAMVL GLVAGLISIG GAKCLPVCCN RVL GIHHIS VMHSIFSLLG LLGEITYIVL LVLHTVWNGN GMIGFQVLLS IGELSLAIVI ALTSGLLTGL LLNLKIWKAP HVAK YFDDQ VFWKFPHLAV GF UniProtKB: Blood group Rh(CE) polypeptide |
-Macromolecule #2: Ammonium transporter Rh type A
| Macromolecule | Name: Ammonium transporter Rh type A / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 44.229629 KDa |
| Sequence | String: MRFTFPLMAI VLEIAMIVLF GLFVEYETDQ TVLEQLNITK PTDMGIFFEL YPLFQDVHVM IFVGFGFLMT FLKKYGFSSV GINLLVAAL GLQWGTIVQG ILQSQGQKFN IGIKNMINAD FSAATVLISF GAVLGKTSPT QMLIMTILEI VFFAHNEYLV S EIFKASDI ...String: MRFTFPLMAI VLEIAMIVLF GLFVEYETDQ TVLEQLNITK PTDMGIFFEL YPLFQDVHVM IFVGFGFLMT FLKKYGFSSV GINLLVAAL GLQWGTIVQG ILQSQGQKFN IGIKNMINAD FSAATVLISF GAVLGKTSPT QMLIMTILEI VFFAHNEYLV S EIFKASDI GASMTIHAFG AYFGLAVAGI LYRSGLRKGH ENEESAYYSD LFAMIGTLFL WMFWPSFNSA IAEPGDKQCR AI VNTYFSL AACVLTAFAF SSLVEHRGKL NMVHIQNATL AGGVAVGTCA DMAIHPFGSM IIGSIAGMVS VLGYKFLTPL FTT KLRIHD TCGVHNLHGL PGVVGGLAGI VAVAMGASNT SMAMQAAALG SSIGTAVVGG LMTGLILKLP LWGQPSDQNC YDDS VYWKV PKTR UniProtKB: Ammonium transporter Rh type A |
-Macromolecule #3: Ankyrin-1
| Macromolecule | Name: Ankyrin-1 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 206.522625 KDa |
| Sequence | String: MPYSVGFREA DAATSFLRAA RSGNLDKALD HLRNGVDINT CNQNGLNGLH LASKEGHVKM VVELLHKEII LETTTKKGNT ALHIAALAG QDEVVRELVN YGANVNAQSQ KGFTPLYMAA QENHLEVVKF LLENGANQNV ATEDGFTPLA VALQQGHENV V AHLINYGT ...String: MPYSVGFREA DAATSFLRAA RSGNLDKALD HLRNGVDINT CNQNGLNGLH LASKEGHVKM VVELLHKEII LETTTKKGNT ALHIAALAG QDEVVRELVN YGANVNAQSQ KGFTPLYMAA QENHLEVVKF LLENGANQNV ATEDGFTPLA VALQQGHENV V AHLINYGT KGKVRLPALH IAARNDDTRT AAVLLQNDPN PDVLSKTGFT PLHIAAHYEN LNVAQLLLNR GASVNFTPQN GI TPLHIAS RRGNVIMVRL LLDRGAQIET KTKDELTPLH CAARNGHVRI SEILLDHGAP IQAKTKNGLS PIHMAAQGDH LDC VRLLLQ YDAEIDDITL DHLTPLHVAA HCGHHRVAKV LLDKGAKPNS RALNGFTPLH IACKKNHVRV MELLLKTGAS IDAV TESGL TPLHVASFMG HLPIVKNLLQ RGASPNVSNV KVETPLHMAA RAGHTEVAKY LLQNKAKVNA KAKDDQTPLH CAARI GHTN MVKLLLENNA NPNLATTAGH TPLHIAAREG HVETVLALLE KEASQACMTK KGFTPLHVAA KYGKVRVAEL LLERDA HPN AAGKNGLTPL HVAVHHNNLD IVKLLLPRGG SPHSPAWNGY TPLHIAAKQN QVEVARSLLQ YGGSANAESV QGVTPLH LA AQEGHAEMVA LLLSKQANGN LGNKSGLTPL HLVAQEGHVP VADVLIKHGV MVDATTRMGY TPLHVASHYG NIKLVKFL L QHQADVNAKT KLGYSPLHQA AQQGHTDIVT LLLKNGASPN EVSSDGTTPL AIAKRLGYIS VTDVLKVVTD ETSFVLVSD KHRMSFPETV DEILDVSEDE GEELISFKAE RRDSRDVDEE KELLDFVPKL DQVVESPAIP RIPCAMPETV VIRSEEQEQA SKEYDEDSL IPSSPATETS DNISPVASPV HTGFLVSFMV DARGGSMRGS RHNGLRVVIP PRTCAAPTRI TCRLVKPQKL S TPPPLAEE EGLASRIIAL GPTGAQFLSP VIVEIPHFAS HGRGDRELVV LRSENGSVWK EHRSRYGESY LDQILNGMDE EL GSLEELE KKRVCRIITT DFPLYFVIMS RLCQDYDTIG PEGGSLKSKL VPLVQATFPE NAVTKRVKLA LQAQPVPDEL VTK LLGNQA TFSPIVTVEP RRRKFHRPIG LRIPLPPSWT DNPRDSGEGD TTSLRLLCSV IGGTDQAQWE DITGTTKLVY ANEC ANFTT NVSARFWLSD CPRTAEAVNF ATLLYKELTA VPYMAKFVIF AKMNDPREGR LRCYCMTDDK VDKTLEQHEN FVEVA RSRD IEVLEGMSLF AELSGNLVPV KKAAQQRSFH FQSFRENRLA MPVKVRDSSR EPGGSLSFLR KAMKYEDTQH ILCHLN ITM PPCAKGSGAE DRRRTPTPLA LRYSILSEST PGSLSGTEQA EMKMAVISEH LGLSWAELAR ELQFSVEDIN RIRVENP NS LLEQSVALLN LWVIREGQNA NMENLYTALQ SIDRGEIVNM LEGSGRQSRN LKPDRRHTDR DYSLSPSQMN GYSSLQDE L LSPASLGCAL SSPLRADQYW NEVAVLDAIP LAATEHDTML EMSDMQVWSA GLTPSLVTAE DSSLECSKAE DSDATGHEW KLEGALSEEP RGPELGSLEL VEDDTVDSDA TNGLIDLLEQ EEGQRSEEKL PGSKRQDDAT GAGQDSENEV SLVSGHQRGQ ARITHSPTV SQVTERSQDR LQDWDADGSI VSYLQDAAQG SWQEEVTQGP HSFQGTSTMT EGLEPGGSQE YEKVLVSVSE H TWTEQPEA ESSQADRDRR QQGQEEQVQE AKNTFTQVVQ GNEFQNIPGE QVTEEQFTDE QGNIVTKKII RKVVRQIDLS SA DAAQEHE EVTVEGPLED PSELEVDIDY FMKHSKDHTS TPNP UniProtKB: Ankyrin-1 |
-Macromolecule #4: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 4 / Number of copies: 2 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #5: Digitonin
| Macromolecule | Name: Digitonin / type: ligand / ID: 5 / Number of copies: 1 / Formula: AJP |
|---|---|
| Molecular weight | Theoretical: 1.229312 KDa |
| Chemical component information |  ChemComp-AJP: |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 178 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 8 mg/mL |
|---|---|
| Buffer | pH: 7.4 Details: Final gel filtration buffer contained 0.05% w/v digitonin, 130 mM KCl, 20 mM HEPES, pH 7.4, 1 mM ATP, 1 mM MgCl2, 1 mM PMSF. Peak fractions were concentrated to 8 mg/mL, and 0.01% w/v ...Details: Final gel filtration buffer contained 0.05% w/v digitonin, 130 mM KCl, 20 mM HEPES, pH 7.4, 1 mM ATP, 1 mM MgCl2, 1 mM PMSF. Peak fractions were concentrated to 8 mg/mL, and 0.01% w/v glycyrrhizic acid was added immediately prior to vitrification. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: 4-6 seconds, wait time 30 seconds. |
| Details | Ankyrin complex mixture purified from digitonin-solubilized erythrocyte ghost membranes |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 2 / Number real images: 14464 / Average exposure time: 2.5 sec. / Average electron dose: 58.0 e/Å2 / Details: Two grids were imaged in a single session. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)