[English] 日本語
 Yorodumi
Yorodumi- EMDB-26734: Cryo-EM structure of WAVE regulatory complex with Rac1 bound on b... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 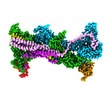 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
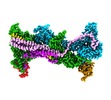
| Title | Cryo-EM structure of WAVE regulatory complex with Rac1 bound on both A and D site | |||||||||
 Map data Map data | WRC230VCA-2Rac1 complex sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | actin regulator / GTPase binding protein / cytoskeletal regulator / CELL INVASION | |||||||||
| Function / homology |  Function and homology information Function and homology informationperipheral region of growth cone / negative regulation of synaptic vesicle recycling / SCAR complex / positive regulation of neurotrophin TRK receptor signaling pathway / lamellipodium morphogenesis / positive regulation of Arp2/3 complex-mediated actin nucleation / Arp2/3 complex binding / modification of synaptic structure / regulation of actin polymerization or depolymerization / central region of growth cone ...peripheral region of growth cone / negative regulation of synaptic vesicle recycling / SCAR complex / positive regulation of neurotrophin TRK receptor signaling pathway / lamellipodium morphogenesis / positive regulation of Arp2/3 complex-mediated actin nucleation / Arp2/3 complex binding / modification of synaptic structure / regulation of actin polymerization or depolymerization / central region of growth cone / modification of postsynaptic actin cytoskeleton / regulation of respiratory burst / dendrite extension / negative regulation of interleukin-23 production / regulation of neutrophil migration / localization within membrane / regulation of translation at postsynapse, modulating synaptic transmission / Activated NTRK2 signals through CDK5 / filopodium tip / negative regulation of receptor-mediated endocytosis / regulation of hydrogen peroxide metabolic process / ruffle assembly / NTRK2 activates RAC1 / Inactivation of CDC42 and RAC1 / NADPH oxidase complex / engulfment of apoptotic cell / WNT5:FZD7-mediated leishmania damping / respiratory burst / regulation of modification of postsynaptic actin cytoskeleton / regulation of actin filament polymerization / SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion / cortical cytoskeleton organization / hepatocyte growth factor receptor signaling pathway / RNA 7-methylguanosine cap binding / ruffle organization / thioesterase binding / regulation of stress fiber assembly / negative regulation of fibroblast migration / cell projection assembly / RHO GTPases activate CIT / axon extension / sphingosine-1-phosphate receptor signaling pathway / Nef and signal transduction / PCP/CE pathway / positive regulation of neutrophil chemotaxis / regulation of nitric oxide biosynthetic process / RHO GTPases activate KTN1 / Activation of RAC1 / motor neuron axon guidance / positive regulation of ruffle assembly / regulation of lamellipodium assembly / positive regulation of dendrite development / Azathioprine ADME / MET activates RAP1 and RAC1 / DCC mediated attractive signaling / positive regulation of cell-substrate adhesion / Wnt signaling pathway, planar cell polarity pathway / Sema4D mediated inhibition of cell attachment and migration / CD28 dependent Vav1 pathway / Ephrin signaling / regulation of myelination / protein kinase A binding / lamellipodium assembly / cortical actin cytoskeleton organization / regulation of cell size / establishment or maintenance of cell polarity / DSCAM interactions / Activation of RAC1 downstream of NMDARs / small GTPase-mediated signal transduction / Rho GDP-dissociation inhibitor binding / positive regulation of Rho protein signal transduction / positive regulation of actin filament polymerization / NRAGE signals death through JNK / Rac protein signal transduction / protein kinase A regulatory subunit binding / dendritic growth cone / filamentous actin / RHO GTPases activate PAKs / positive regulation of focal adhesion assembly / lamellipodium membrane / semaphorin-plexin signaling pathway / Sema3A PAK dependent Axon repulsion / ficolin-1-rich granule membrane / excitatory synapse / RHOG GTPase cycle / RHO GTPases Activate NADPH Oxidases / EPH-ephrin mediated repulsion of cells / RHO GTPases Activate WASPs and WAVEs / RAC2 GTPase cycle / localization / RAC3 GTPase cycle / RHO GTPases activate IQGAPs / anatomical structure morphogenesis / positive regulation of axon extension / axonal growth cone / PTK6 Regulates RHO GTPases, RAS GTPase and MAP kinases / positive regulation of lamellipodium assembly / signaling adaptor activity / response to electrical stimulus / positive regulation of substrate adhesion-dependent cell spreading Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
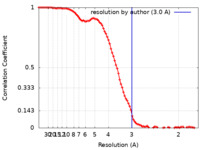
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Ding B / Yang S / Chen B / Chowdhury S | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structures reveal a key mechanism of WAVE regulatory complex activation by Rac1 GTPase. Authors: Bojian Ding / Sheng Yang / Matthias Schaks / Yijun Liu / Abbigale J Brown / Klemens Rottner / Saikat Chowdhury / Baoyu Chen /    Abstract: The Rho-family GTPase Rac1 activates the WAVE regulatory complex (WRC) to drive Arp2/3 complex-mediated actin polymerization in many essential processes. Rac1 binds to WRC at two distinct sites-the ...The Rho-family GTPase Rac1 activates the WAVE regulatory complex (WRC) to drive Arp2/3 complex-mediated actin polymerization in many essential processes. Rac1 binds to WRC at two distinct sites-the A and D sites. Precisely how Rac1 binds and how the binding triggers WRC activation remain unknown. Here we report WRC structures by itself, and when bound to single or double Rac1 molecules, at ~3 Å resolutions by cryogenic-electron microscopy. The structures reveal that Rac1 binds to the two sites by distinct mechanisms, and binding to the A site, but not the D site, drives WRC activation. Activation involves a series of unique conformational changes leading to the release of sequestered WCA (WH2-central-acidic) polypeptide, which stimulates the Arp2/3 complex to polymerize actin. Together with biochemical and cellular analyses, the structures provide a novel mechanistic understanding of how the Rac1-WRC-Arp2/3-actin signaling axis is regulated in diverse biological processes and diseases. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26734.map.gz emd_26734.map.gz | 13.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26734-v30.xml emd-26734-v30.xml emd-26734.xml emd-26734.xml | 32.6 KB 32.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26734_fsc.xml emd_26734_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_26734.png emd_26734.png | 142 KB | ||
| Masks |  emd_26734_msk_1.map emd_26734_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-26734.cif.gz emd-26734.cif.gz | 9.3 KB | ||
| Others |  emd_26734_additional_1.map.gz emd_26734_additional_1.map.gz emd_26734_half_map_1.map.gz emd_26734_half_map_1.map.gz emd_26734_half_map_2.map.gz emd_26734_half_map_2.map.gz | 48.1 MB 46.7 MB 46.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26734 http://ftp.pdbj.org/pub/emdb/structures/EMD-26734 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26734 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26734 | HTTPS FTP |
-Validation report
| Summary document |  emd_26734_validation.pdf.gz emd_26734_validation.pdf.gz | 903.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_26734_full_validation.pdf.gz emd_26734_full_validation.pdf.gz | 903.1 KB | Display | |
| Data in XML |  emd_26734_validation.xml.gz emd_26734_validation.xml.gz | 16.3 KB | Display | |
| Data in CIF |  emd_26734_validation.cif.gz emd_26734_validation.cif.gz | 21.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26734 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26734 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26734 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26734 | HTTPS FTP |
-Related structure data
| Related structure data |  7useMC  7uscC  7usdC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26734.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26734.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | WRC230VCA-2Rac1 complex sharpened map | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8757 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_26734_msk_1.map emd_26734_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: WRC230VCA-2Rac1 complex masked unsharpened map
| File | emd_26734_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | WRC230VCA-2Rac1 complex masked unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: WRC230VCA-2Rac1 complex unfiltered unmasked unsharpened first half map...
| File | emd_26734_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | WRC230VCA-2Rac1 complex unfiltered unmasked unsharpened first half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: WRC230VCA-2Rac1 complex unfiltered unmasked unsharpened second half map...
| File | emd_26734_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | WRC230VCA-2Rac1 complex unfiltered unmasked unsharpened second half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : WAVE regulatory complex with Rac1 bound to A and D sites
+Supramolecule #1: WAVE regulatory complex with Rac1 bound to A and D sites
+Macromolecule #1: Cytoplasmic FMR1-interacting protein 1
+Macromolecule #2: Nck-associated protein 1
+Macromolecule #3: Wiskott-Aldrich syndrome protein family member 1
+Macromolecule #4: Protein BRICK1
+Macromolecule #5: Abl interactor 2
+Macromolecule #6: Ras-related C3 botulinum toxin substrate 1
+Macromolecule #7: Ras-related C3 botulinum toxin substrate 1
+Macromolecule #8: PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER
+Macromolecule #9: MAGNESIUM ION
+Macromolecule #10: GUANOSINE-5'-TRIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.15 mg/mL |
|---|---|
| Buffer | pH: 7 |
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 120 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.019 kPa / Details: 15 mA |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277.15 K / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Details | Data were collected by shifting the stage to target exposure positions. |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 1285 / Average exposure time: 40.0 sec. / Average electron dose: 41.34 e/Å2 Details: Each micrograph was acquired as dose-fractionated movies consisting of 62 frames per movie. |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.2 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 120000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






















 Z
Z Y
Y X
X


























 Trichoplusia ni (cabbage looper)
Trichoplusia ni (cabbage looper)