+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structure of lineage I (Pinneo) Lassa virus glycoprotein bound to Fab 25.10C | ||||||||||||
 マップデータ マップデータ | Sharpened map | ||||||||||||
 試料 試料 |
| ||||||||||||
 キーワード キーワード | viral glycoprotein / antibody Fab fragment / VIRAL PROTEIN / VIRAL PROTEIN-IMMUNE SYSTEM complex | ||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報host cell Golgi membrane / receptor-mediated endocytosis of virus by host cell / host cell endoplasmic reticulum membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / membrane / metal ion binding 類似検索 - 分子機能 | ||||||||||||
| 生物種 |  Homo sapiens (ヒト) / Homo sapiens (ヒト) /  Lassa mammarenavirus (ウイルス) Lassa mammarenavirus (ウイルス) | ||||||||||||
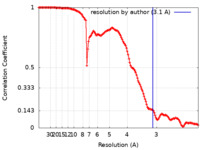
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.1 Å | ||||||||||||
 データ登録者 データ登録者 | Buck TK / Enriquez AE / Hastie KM | ||||||||||||
| 資金援助 |  米国, 3件 米国, 3件
| ||||||||||||
 引用 引用 |  ジャーナル: mBio / 年: 2022 ジャーナル: mBio / 年: 2022タイトル: Neutralizing Antibodies against Lassa Virus Lineage I. 著者: Tierra K Buck / Adrian S Enriquez / Sharon L Schendel / Michelle A Zandonatti / Stephanie S Harkins / Haoyang Li / Alex Moon-Walker / James E Robinson / Luis M Branco / Robert F Garry / Erica ...著者: Tierra K Buck / Adrian S Enriquez / Sharon L Schendel / Michelle A Zandonatti / Stephanie S Harkins / Haoyang Li / Alex Moon-Walker / James E Robinson / Luis M Branco / Robert F Garry / Erica Ollmann Saphire / Kathryn M Hastie /  要旨: Lassa virus (LASV) is the causative agent of the deadly Lassa fever (LF). Seven distinct LASV lineages circulate through western Africa, among which lineage I (LI), the first to be identified, is ...Lassa virus (LASV) is the causative agent of the deadly Lassa fever (LF). Seven distinct LASV lineages circulate through western Africa, among which lineage I (LI), the first to be identified, is particularly resistant to antibody neutralization. Lineage I LASV evades neutralization by half of known antibodies in the GPC-A antibody competition group and all but one of the antibodies in the GPC-B competition group. Here, we solve two cryo-electron microscopy (cryo-EM) structures of LI GP in complex with a GPC-A and a GPC-B antibody. We used complementary structural and biochemical techniques to identify single-amino-acid substitutions in LI that are responsible for immune evasion by each antibody group. Further, we show that LI infection is more dependent on the endosomal receptor lysosome-associated membrane protein 1 (LAMP1) for viral entry relative to LIV. In the absence of LAMP1, LI requires a more acidic fusion pH to initiate membrane fusion with the host cell relative to LIV. No vaccine or therapeutics are approved to prevent LASV infection or treat LF. All vaccine platforms currently under development present only the LIV GP sequence. However, our data suggest that the high genetic diversity of LASV may be problematic for designing both a broadly reactive immunogen and therapeutic. Here, we examine antibodies that are highly potent against LIV yet are ineffective against LI. By pinpointing LI mutations responsible for this decrease in antibody efficacy, we suggest that future vaccine platforms may need to incorporate specific LI-like mutations in order to generate a broadly neutralizing antibody response against all LASV lineages. | ||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_26458.map.gz emd_26458.map.gz | 191.9 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-26458-v30.xml emd-26458-v30.xml emd-26458.xml emd-26458.xml | 21.3 KB 21.3 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_26458_fsc.xml emd_26458_fsc.xml | 13.3 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_26458.png emd_26458.png | 43 KB | ||
| Filedesc metadata |  emd-26458.cif.gz emd-26458.cif.gz | 6.4 KB | ||
| その他 |  emd_26458_additional_1.map.gz emd_26458_additional_1.map.gz emd_26458_half_map_1.map.gz emd_26458_half_map_1.map.gz emd_26458_half_map_2.map.gz emd_26458_half_map_2.map.gz | 108.3 MB 200.1 MB 200.1 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26458 http://ftp.pdbj.org/pub/emdb/structures/EMD-26458 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26458 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26458 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  7udsMC  7ul7C M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_26458.map.gz / 形式: CCP4 / 大きさ: 216 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_26458.map.gz / 形式: CCP4 / 大きさ: 216 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Sharpened map | ||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.096 Å | ||||||||||||||||||||||||||||||||||||
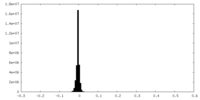
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-追加マップ: Unsharpened map
| ファイル | emd_26458_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Unsharpened map | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
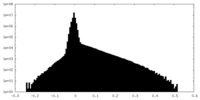
| 密度ヒストグラム |
-ハーフマップ: Half map A
| ファイル | emd_26458_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Half map A | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: Half map B
| ファイル | emd_26458_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Half map B | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Lineage I Lassa virus glycoprotein bound to Fab 25.10C
| 全体 | 名称: Lineage I Lassa virus glycoprotein bound to Fab 25.10C |
|---|---|
| 要素 |
|
-超分子 #1: Lineage I Lassa virus glycoprotein bound to Fab 25.10C
| 超分子 | 名称: Lineage I Lassa virus glycoprotein bound to Fab 25.10C タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1-#4 |
|---|---|
| 分子量 | 理論値: 350 KDa |
-分子 #1: 25.10C Fab Heavy Chain
| 分子 | 名称: 25.10C Fab Heavy Chain / タイプ: protein_or_peptide / ID: 1 / コピー数: 3 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 24.27025 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: QVQLQESGGG LVKPGGSLRL SCTASGFNFN KYNMNWVRQA PGKGLEWVSS ISALSTYIYY ADSLKGRFTV SRDNAKNSLF LQMNSLRDD DTAVYYCARE IRRASTWSAD LWGRGTLVTV SSASTKGPSV FPLAPSSKST SGGTAALGCL VKDYFPEPVT V SWNSGALT ...文字列: QVQLQESGGG LVKPGGSLRL SCTASGFNFN KYNMNWVRQA PGKGLEWVSS ISALSTYIYY ADSLKGRFTV SRDNAKNSLF LQMNSLRDD DTAVYYCARE IRRASTWSAD LWGRGTLVTV SSASTKGPSV FPLAPSSKST SGGTAALGCL VKDYFPEPVT V SWNSGALT SGVHTFPAVL QSSGLYSLSS VVTVPSSSLG TQTYICNVNH KPSNTKVDKR VEPKSCDK |
-分子 #2: 25.10C Fab Light Chain
| 分子 | 名称: 25.10C Fab Light Chain / タイプ: protein_or_peptide / ID: 2 / コピー数: 3 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 22.812377 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: IQMTQSPSSL SASVGDRVII TCRASQSISS SLNWYQQKPG KAPKLLIYAA VNLETGVPSR FSGSGFGTDF TLAISNVQPE DFATYYCQQ SDTRTFGRGT KLDVKRTVAA PSVFIFPPSD EQLKSGTASV VCLLNNFYPR EAKVQWKVDN ALQSGNSQES V TEQDSKDS ...文字列: IQMTQSPSSL SASVGDRVII TCRASQSISS SLNWYQQKPG KAPKLLIYAA VNLETGVPSR FSGSGFGTDF TLAISNVQPE DFATYYCQQ SDTRTFGRGT KLDVKRTVAA PSVFIFPPSD EQLKSGTASV VCLLNNFYPR EAKVQWKVDN ALQSGNSQES V TEQDSKDS TYSLSSTLTL SKADYEKHKV YACEVTHQGL SSPVTKSFNR G |
-分子 #3: Glycoprotein G1
| 分子 | 名称: Glycoprotein G1 / タイプ: protein_or_peptide / ID: 3 / コピー数: 3 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Lassa mammarenavirus (ウイルス) Lassa mammarenavirus (ウイルス) |
| 分子量 | 理論値: 28.987398 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MGQIITFFQE VPHVIEEVMN IVLIALSLLA ILKGLYNIAT CGIIGLVAFL FLCGKSCSLT LKGGYELQTL ELNMETLNMT MPLSCTKNS SHHYIRVGNE TGLELTLTNT SIINHKFCNL SDAHKKNLYD HALMSIISTF HLSIPNFNQY EAMSCDFNGG K ISVQYNLS ...文字列: MGQIITFFQE VPHVIEEVMN IVLIALSLLA ILKGLYNIAT CGIIGLVAFL FLCGKSCSLT LKGGYELQTL ELNMETLNMT MPLSCTKNS SHHYIRVGNE TGLELTLTNT SIINHKFCNL SDAHKKNLYD HALMSIISTF HLSIPNFNQY EAMSCDFNGG K ISVQYNLS HSYAGDAAEH CGTVANGVLQ TFMRMAWGGR YIALDSGCGN WDCIMTSYQY LIIQNTTWED HCQFSRPSPI GY LGLLSQR TRDIYISRRR R UniProtKB: Pre-glycoprotein polyprotein GP complex |
-分子 #4: Glycoprotein G2
| 分子 | 名称: Glycoprotein G2 / タイプ: protein_or_peptide / ID: 4 / コピー数: 3 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Lassa mammarenavirus (ウイルス) Lassa mammarenavirus (ウイルス) |
| 分子量 | 理論値: 23.348145 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: GTFTWTLSDS EGNETPGGYC LTRWMLIEAE LKCFGNTAVA KCNEKHDEEF CDMLRLFDFN KQAIRRLKAP AQMSIQLINK AVNALINDQ LIMKNHLRDI MCIPYCNYSK YWYLNHTSSG RTSLPKCWLI SNGSYLNETQ FSDDIEQQAD NMITEMLQKE Y LPETGLVD ...文字列: GTFTWTLSDS EGNETPGGYC LTRWMLIEAE LKCFGNTAVA KCNEKHDEEF CDMLRLFDFN KQAIRRLKAP AQMSIQLINK AVNALINDQ LIMKNHLRDI MCIPYCNYSK YWYLNHTSSG RTSLPKCWLI SNGSYLNETQ FSDDIEQQAD NMITEMLQKE Y LPETGLVD LEVDDDDKAG WSHPQFEKGG GSGGGSGGGS WSHPQFEK UniProtKB: Pre-glycoprotein polyprotein GP complex |
-分子 #8: 2-acetamido-2-deoxy-beta-D-glucopyranose
| 分子 | 名称: 2-acetamido-2-deoxy-beta-D-glucopyranose / タイプ: ligand / ID: 8 / コピー数: 10 / 式: NAG |
|---|---|
| 分子量 | 理論値: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 構成要素:
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 298 K |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: SUPER-RESOLUTION / 平均電子線量: 55.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.5 µm / 最小 デフォーカス(公称値): 1.0 µm |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)