[English] 日本語
 Yorodumi
Yorodumi- EMDB-26309: Cryo-EM structure of the pancreatic ATP-sensitive potassium chann... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the pancreatic ATP-sensitive potassium channel bound to ATP and in the presence of carbamazepine with Kir6.2-CTD in the up conformation | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | KATP channel / SUR1 / Kir6.2 / carbamazepine / CBM / CBZ / sulfonylurea receptor / potassium transport / MEMBRANE PROTEIN / metabolic sensor / trafficking defects | |||||||||
| Function / homology |  Function and homology information Function and homology informationRegulation of insulin secretion / ATP sensitive Potassium channels / ABC-family proteins mediated transport / response to resveratrol / ATP-activated inward rectifier potassium channel activity / inward rectifying potassium channel / cell body fiber / sulfonylurea receptor activity / ventricular cardiac muscle tissue development / CAMKK-AMPK signaling cascade ...Regulation of insulin secretion / ATP sensitive Potassium channels / ABC-family proteins mediated transport / response to resveratrol / ATP-activated inward rectifier potassium channel activity / inward rectifying potassium channel / cell body fiber / sulfonylurea receptor activity / ventricular cardiac muscle tissue development / CAMKK-AMPK signaling cascade / voltage-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / inward rectifier potassium channel activity / ATPase-coupled monoatomic cation transmembrane transporter activity / nervous system process / regulation of monoatomic ion transmembrane transport / inorganic cation transmembrane transport / ankyrin binding / Ion homeostasis / response to ATP / response to testosterone / voltage-gated potassium channel activity / potassium ion import across plasma membrane / action potential / regulation of insulin secretion / potassium channel activity / intercalated disc / axolemma / ABC-type transporter activity / negative regulation of insulin secretion / potassium ion transmembrane transport / T-tubule / heat shock protein binding / positive regulation of protein localization to plasma membrane / potassium ion transport / acrosomal vesicle / regulation of membrane potential / response to ischemia / determination of adult lifespan / cellular response to glucose stimulus / sarcolemma / cellular response to nicotine / glucose metabolic process / nuclear envelope / response to estradiol / cellular response to tumor necrosis factor / presynaptic membrane / transmembrane transporter binding / endosome / response to hypoxia / response to xenobiotic stimulus / neuronal cell body / glutamatergic synapse / apoptotic process / ATP hydrolysis activity / ATP binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Cricetus cricetus (black-bellied hamster) / Cricetus cricetus (black-bellied hamster) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 5.2 Å | |||||||||
 Authors Authors | Shyng SL / Sung MW / Driggers CM | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2019 Journal: Elife / Year: 2019Title: Mechanism of pharmacochaperoning in a mammalian K channel revealed by cryo-EM. Authors: Gregory M Martin / Min Woo Sung / Zhongying Yang / Laura M Innes / Balamurugan Kandasamy / Larry L David / Craig Yoshioka / Show-Ling Shyng /  Abstract: ATP-sensitive potassium (K) channels composed of a pore-forming Kir6.2 potassium channel and a regulatory ABC transporter sulfonylurea receptor 1 (SUR1) regulate insulin secretion in pancreatic β- ...ATP-sensitive potassium (K) channels composed of a pore-forming Kir6.2 potassium channel and a regulatory ABC transporter sulfonylurea receptor 1 (SUR1) regulate insulin secretion in pancreatic β-cells to maintain glucose homeostasis. Mutations that impair channel folding or assembly prevent cell surface expression and cause congenital hyperinsulinism. Structurally diverse K inhibitors are known to act as pharmacochaperones to correct mutant channel expression, but the mechanism is unknown. Here, we compare cryoEM structures of a mammalian K channel bound to pharmacochaperones glibenclamide, repaglinide, and carbamazepine. We found all three drugs bind within a common pocket in SUR1. Further, we found the N-terminus of Kir6.2 inserted within the central cavity of the SUR1 ABC core, adjacent the drug binding pocket. The findings reveal a common mechanism by which diverse compounds stabilize the Kir6.2 N-terminus within SUR1's ABC core, allowing it to act as a firm 'handle' for the assembly of metastable mutant SUR1-Kir6.2 complexes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26309.map.gz emd_26309.map.gz | 2.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26309-v30.xml emd-26309-v30.xml emd-26309.xml emd-26309.xml | 16.5 KB 16.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_26309.png emd_26309.png | 116.9 KB | ||
| Filedesc metadata |  emd-26309.cif.gz emd-26309.cif.gz | 6.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26309 http://ftp.pdbj.org/pub/emdb/structures/EMD-26309 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26309 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26309 | HTTPS FTP |
-Related structure data
| Related structure data |  7u7mMC  7tysC  7tytC  7u1eC  7u1qC  7u1sC  7u24C  7u2xC  7u6yC  7uaaC  7uqrC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26309.map.gz / Format: CCP4 / Size: 2.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26309.map.gz / Format: CCP4 / Size: 2.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.71 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : KATP-CBZ-ATP-CTDup
| Entire | Name: KATP-CBZ-ATP-CTDup |
|---|---|
| Components |
|
-Supramolecule #1: KATP-CBZ-ATP-CTDup
| Supramolecule | Name: KATP-CBZ-ATP-CTDup / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Cricetus cricetus (black-bellied hamster) Cricetus cricetus (black-bellied hamster) |
-Supramolecule #2: Kir6.2
| Supramolecule | Name: Kir6.2 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 / Details: Adenovirus-based infection of INS-1 cells |
|---|---|
| Source (natural) | Organism:  Cricetus cricetus (black-bellied hamster) Cricetus cricetus (black-bellied hamster) |
-Supramolecule #3: SUR1
| Supramolecule | Name: SUR1 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 / Details: Adenovirus-based infection of INS-1 cells |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: ATP-sensitive inward rectifier potassium channel 11
| Macromolecule | Name: ATP-sensitive inward rectifier potassium channel 11 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MLSRKGIIPE EYVLTRLAED PTEPRYRTRE RRARFVSKKG NCNVAHKNIR EQGRFLQDVF TTLVDLKWP HTLLIFTMSF LCSWLLFAMV WWLIAFAHGD LAPGEGTNVP CVTSIHSFSS A FLFSIEVQ VTIGFGGRMV TEECPLAILI LIVQNIVGLM INAIMLGCIF ...String: MLSRKGIIPE EYVLTRLAED PTEPRYRTRE RRARFVSKKG NCNVAHKNIR EQGRFLQDVF TTLVDLKWP HTLLIFTMSF LCSWLLFAMV WWLIAFAHGD LAPGEGTNVP CVTSIHSFSS A FLFSIEVQ VTIGFGGRMV TEECPLAILI LIVQNIVGLM INAIMLGCIF MKTAQAHRRA ET LIFSKHA VITLRHGRLC FMLRVGDLRK SMIISATIHM QVVRKTTSPE GEVVPLHQVD IPM ENGVGG NSIFLVAPLI IYHVIDSNSP LYDLAPSDLH HHQDLEIIVI LEGVVETTGI TTQA RTSYL ADEILWGQRF VPIVAEEDGR YSVDYSKFGN TVKVPTPLCT ARQLDEDRSL LDALT LASS RGPLRKRSVA VAKAKPKFSI SPDSLS UniProtKB: ATP-sensitive inward rectifier potassium channel 11 |
-Macromolecule #2: ATP-binding cassette sub-family C member 8
| Macromolecule | Name: ATP-binding cassette sub-family C member 8 / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Cricetus cricetus (black-bellied hamster) Cricetus cricetus (black-bellied hamster) |
| Sequence | String: MPLAFCGTEN HSAAYRVDQG VLNNGCFVDA LNVVPHVFLL FITFPILFIG WGSQSSKVHI HHSTWLHFP GHNLRWILTF ILLFVLVCEI AEGILSDGVT ESRHLHLYMP AGMAFMAAIT S VVYYHNIE TSNFPKLLIA LLIYWTLAFI TKTIKFVKFY DHAIGFSQLR ...String: MPLAFCGTEN HSAAYRVDQG VLNNGCFVDA LNVVPHVFLL FITFPILFIG WGSQSSKVHI HHSTWLHFP GHNLRWILTF ILLFVLVCEI AEGILSDGVT ESRHLHLYMP AGMAFMAAIT S VVYYHNIE TSNFPKLLIA LLIYWTLAFI TKTIKFVKFY DHAIGFSQLR FCLTGLLVIL YG MLLLVEV NVIRVRRYIF FKTPREVKPP EDLQDLGVRF LQPFVNLLSK GTYWWMNAFI KTA HKKPID LRAIAKLPIA MRALTNYQRL CVAFDAQARK DTQSPQGARA IWRALCHAFG RRLI LSSTF RILADLLGFA GPLCIFGIVD HLGKENHVFQ PKTQFLGVYF VSSQEFLGNA YVLAV LLFL ALLLQRTFLQ ASYYVAIETG INLRGAIQTK IYNKIMHMST SNLSMGEMTA GQICNL VAI DTNQLMWFFF LCPNLWTMPV QIIVGVILLY YILGVSALIG AAVIILLAPV QYFVATK LS QAQRTTLEHS NERLKQTNEM LRGMKLLKLY AWESIFCSRV EVTRRKEMTS LRAFAVYT S ISIFMNTAIP IAAVLITFVG HVSFFKESDL SPSVAFASLS LFHILVTPLF LLSSVVRST VKALVSVQKL SEFLSSAEIR EEQCAPREPA PQGQAGKYQA VPLKVVNRKR PAREEVRDLL GPLQRLAPS MDGDADNFCV QIIGGFFTWT PDGIPTLSNI TIRIPRGQLT MIVGQVGCGK S SLLLATLG EMQKVSGAVF WNSNLPDSEG EDPSSPERET AAGSDIRSRG PVAYASQKPW LL NATVEEN ITFESPFNKQ RYKMVIEACS LQPDIDILPH GDQTQIGERG INLSGGQRQR ISV ARALYQ QTNVVFLDDP FSALDVHLSD HLMQAGILEL LRDDKRTVVL VTHKLQYLPH ADWI IAMKD GTIQREGTLK DFQRSECQLF EHWKTLMNRQ DQELEKETVM ERKASEPSQG LPRAM SSRD GLLLDEEEEE EEAAESEEDD NLSSVLHQRA KIPWRACTKY LSSAGILLLS LLVFSQ LLK HMVLVAIDYW LAKWTDSALV LSPAARNCSL SQECDLDQSV YAMVFTLLCS LGIVLCL VT SVTVEWTGLK VAKRLHRSLL NRIILAPMRF FETTPLGSIL NRFSSDCNTI DQHIPSTL E CLSRSTLLCV SALTVISYVT PVFLVALLPL AVVCYFIQKY FRVASRDLQQ LDDTTQLPL VSHFAETVEG LTTIRAFRYE ARFQQKLLEY TDSNNIASLF LTAANRWLEV CMEYIGACVV LIAAATSIS NSLHRELSAG LVGLGLTYAL MVSNYLNWMV RNLADMEIQL GAVKRIHALL K TEAESYEG LLAPSLIPKN WPDQGKIQIQ NLSVRYDSSL KPVLKHVNTL ISPGQKIGIC GR TGSGKSS FSLAFFRMVD MFEGRIIIDG IDIAKLPLHT LRSRLSIILQ DPVLFSGTIR FNL DPEKKC SDSTLWEALE IAQLKLVVKA LPGGLDAIIT EGGENFSQGQ RQLFCLARAF VRKT SIFIM DEATASIDMA TENILQKVVM TAFADRTVVT IAHRVHTILS ADLVMVLKRG AILEF DKPE TLLSQKDSVF ASFVRADK UniProtKB: ATP-binding cassette sub-family C member 8 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.6 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP EMDB ID: |
|---|---|
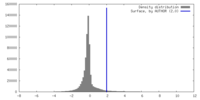
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 5.2 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 50000 |
| Initial angle assignment | Type: COMMON LINE |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
-Atomic model buiding 1
| Initial model |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Output model |  PDB-7u7m: |
 Movie
Movie Controller
Controller

















 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)