[English] 日本語
 Yorodumi
Yorodumi- EMDB-26050: Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta)... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-26050 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
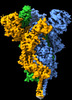
| Title | Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta) in the 1-RBD-up conformation; Subclassification D14 state | |||||||||
 Map data Map data | SARS-CoV-2 variant spike protein (S-GSAS-Delta) in the 1-RBD-up conformation; Subclassification D14 state | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SARS-CoV-2 Spike Protein Trimer / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Gobeil S / Acharya P | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2022 Journal: Mol Cell / Year: 2022Title: Structural diversity of the SARS-CoV-2 Omicron spike. Authors: Sophie M-C Gobeil / Rory Henderson / Victoria Stalls / Katarzyna Janowska / Xiao Huang / Aaron May / Micah Speakman / Esther Beaudoin / Kartik Manne / Dapeng Li / Rob Parks / Maggie Barr / ...Authors: Sophie M-C Gobeil / Rory Henderson / Victoria Stalls / Katarzyna Janowska / Xiao Huang / Aaron May / Micah Speakman / Esther Beaudoin / Kartik Manne / Dapeng Li / Rob Parks / Maggie Barr / Margaret Deyton / Mitchell Martin / Katayoun Mansouri / Robert J Edwards / Amanda Eaton / David C Montefiori / Gregory D Sempowski / Kevin O Saunders / Kevin Wiehe / Wilton Williams / Bette Korber / Barton F Haynes / Priyamvada Acharya /  Abstract: Aided by extensive spike protein mutation, the SARS-CoV-2 Omicron variant overtook the previously dominant Delta variant. Spike conformation plays an essential role in SARS-CoV-2 evolution via ...Aided by extensive spike protein mutation, the SARS-CoV-2 Omicron variant overtook the previously dominant Delta variant. Spike conformation plays an essential role in SARS-CoV-2 evolution via changes in receptor-binding domain (RBD) and neutralizing antibody epitope presentation, affecting virus transmissibility and immune evasion. Here, we determine cryo-EM structures of the Omicron and Delta spikes to understand the conformational impacts of mutations in each. The Omicron spike structure revealed an unusually tightly packed RBD organization with long range impacts that were not observed in the Delta spike. Binding and crystallography revealed increased flexibility at the functionally critical fusion peptide site in the Omicron spike. These results reveal a highly evolved Omicron spike architecture with possible impacts on its high levels of immune evasion and transmissibility. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26050.map.gz emd_26050.map.gz | 96.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26050-v30.xml emd-26050-v30.xml emd-26050.xml emd-26050.xml | 17.2 KB 17.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_26050.png emd_26050.png | 119.6 KB | ||
| Filedesc metadata |  emd-26050.cif.gz emd-26050.cif.gz | 7.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26050 http://ftp.pdbj.org/pub/emdb/structures/EMD-26050 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26050 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26050 | HTTPS FTP |
-Related structure data
| Related structure data |  7tpaMC  7teiC  7tf8C  7thtC  7tl1C  7tl9C  7tlaC  7tlcC  7tldC  7touC  7tovC  7towC  7toxC  7toyC  7tozC  7tp0C  7tp1C  7tp2C  7tp7C  7tp8C  7tp9C  7tpcC  7tpeC  7tpfC  7tphC  7tplC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26050.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26050.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SARS-CoV-2 variant spike protein (S-GSAS-Delta) in the 1-RBD-up conformation; Subclassification D14 state | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta)...
| Entire | Name: Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta) in the 1-RBD-up conformation; Subclassification D14 state |
|---|---|
| Components |
|
-Supramolecule #1: Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta)...
| Supramolecule | Name: Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta) in the 1-RBD-up conformation; Subclassification D14 state type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 142.244328 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLRT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDG VYFASTEKSN IIRGWIFGTT LDSKTQSLLI VNNATNVVIK VCEFQFCNDP FLDVYYHKNN KSWMESGVYS S ANNCTFEY ...String: MFVFLVLLPL VSSQCVNLRT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDG VYFASTEKSN IIRGWIFGTT LDSKTQSLLI VNNATNVVIK VCEFQFCNDP FLDVYYHKNN KSWMESGVYS S ANNCTFEY VSQPFLMDLE GKQGNFKNLR EFVFKNIDGY FKIYSKHTPI NLVRDLPQGF SALEPLVDLP IGINITRFQT LL ALHRSYL TPGDSSSGWT AGAAAYYVGY LQPRTFLLKY NENGTITDAV DCALDPLSET KCTLKSFTVE KGIYQTSNFR VQP TESIVR FPNITNLCPF GEVFNATRFA SVYAWNRKRI SNCVADYSVL YNSASFSTFK CYGVSPTKLN DLCFTNVYAD SFVI RGDEV RQIAPGQTGK IADYNYKLPD DFTGCVIAWN SNNLDSKVGG NYNYRYRLFR KSNLKPFERD ISTEIYQAGS KPCNG VEGF NCYFPLQSYG FQPTNGVGYQ PYRVVVLSFE LLHAPATVCG PKKSTNLVKN KCVNFNFNGL TGTGVLTESN KKFLPF QQF GRDIADTTDA VRDPQTLEIL DITPCSFGGV SVITPGTNTS NQVAVLYQGV NCTEVPVAIH ADQLTPTWRV YSTGSNV FQ TRAGCLIGAE HVNNSYECDI PIGAGICASY QTQTNSRGSA SSVASQSIIA YTMSLGAENS VAYSNNSIAI PTNFTISV T TEILPVSMTK TSVDCTMYIC GDSTECSNLL LQYGSFCTQL NRALTGIAVE QDKNTQEVFA QVKQIYKTPP IKDFGGFNF SQILPDPSKP SKRSFIEDLL FNKVTLADAG FIKQYGDCLG DIAARDLICA QKFNGLTVLP PLLTDEMIAQ YTSALLAGTI TSGWTFGAG AALQIPFAMQ MAYRFNGIGV TQNVLYENQK LIANQFNSAI GKIQDSLSST ASALGKLQNV VNQNAQALNT L VKQLSSNF GAISSVLNDI LSRLDKVEAE VQIDRLITGR LQSLQTYVTQ QLIRAAEIRA SANLAATKMS ECVLGQSKRV DF CGKGYHL MSFPQSAPHG VVFLHVTYVP AQEKNFTTAP AICHDGKAHF PREGVFVSNG THWFVTQRNF YEPQIITTDN TFV SGNCDV VIGIVNNTVY DPLQPELDSF KEELDKYFKN HTSPDVDLGD ISGINASVVN IQKEIDRLNE VAKNLNESLI DLQE LGKYE QGSGYIPEAP RDGQAYVRKD GEWVLLSTFL GRSLEVLFQG PGHHHHHHHH SAWSHPQFEK GGGSGGGGSG GSAWS HPQF EK UniProtKB: Spike glycoprotein |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 34 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 55.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.75 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















