[English] 日本語
 Yorodumi
Yorodumi- EMDB-25740: Composite map of Upright Munc13-1 C1-C2B-MUN-C2C molecule spannin... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-25740 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Composite map of Upright Munc13-1 C1-C2B-MUN-C2C molecule spanning two lipid bilayers | |||||||||
 Map data Map data | Composite map created in UCSF Chimera by combining two maps from independent, focused 3D classifications using the "vop maximum" command | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Synaptic Transmission / Munc13 / Membrane Fusion / EXOCYTOSIS | |||||||||
| Function / homology |  Function and homology information Function and homology informationdense core granule priming / neuronal dense core vesicle exocytosis / diacylglycerol binding / presynaptic dense core vesicle exocytosis / synaptic vesicle docking / regulation of synaptic vesicle priming / synaptic vesicle maturation / positive regulation of glutamate receptor signaling pathway / presynaptic active zone cytoplasmic component / positive regulation of synaptic plasticity ...dense core granule priming / neuronal dense core vesicle exocytosis / diacylglycerol binding / presynaptic dense core vesicle exocytosis / synaptic vesicle docking / regulation of synaptic vesicle priming / synaptic vesicle maturation / positive regulation of glutamate receptor signaling pathway / presynaptic active zone cytoplasmic component / positive regulation of synaptic plasticity / innervation / positive regulation of dendrite extension / neurotransmitter secretion / regulation of short-term neuronal synaptic plasticity / regulation of amyloid precursor protein catabolic process / syntaxin binding / syntaxin-1 binding / positive regulation of neurotransmitter secretion / synaptic vesicle priming / Golgi-associated vesicle / neuromuscular junction development / spectrin binding / presynaptic active zone / synaptic vesicle exocytosis / excitatory synapse / calyx of Held / amyloid-beta metabolic process / SNARE binding / synaptic transmission, glutamatergic / synaptic membrane / long-term synaptic potentiation / neuromuscular junction / terminal bouton / phospholipid binding / synaptic vesicle membrane / presynapse / presynaptic membrane / cell differentiation / calmodulin binding / neuron projection / protein domain specific binding / axon / glutamatergic synapse / calcium ion binding / synapse / protein-containing complex binding / protein-containing complex / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 10.0 Å | |||||||||
 Authors Authors | Grushin K / Sindelar CV | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: Munc13 structural transitions and oligomers that may choreograph successive stages in vesicle priming for neurotransmitter release. Authors: Kirill Grushin / R Venkat Kalyana Sundaram / Charles V Sindelar / James E Rothman /  Abstract: How can exactly six SNARE complexes be assembled under each synaptic vesicle? Here we report cryo-EM crystal structures of the core domain of Munc13, the key chaperone that initiates SNAREpin ...How can exactly six SNARE complexes be assembled under each synaptic vesicle? Here we report cryo-EM crystal structures of the core domain of Munc13, the key chaperone that initiates SNAREpin assembly. The functional core of Munc13, consisting of C1-C2B-MUN-C2C (Munc13C) spontaneously crystallizes between phosphatidylserine-rich bilayers in two distinct conformations, each in a radically different oligomeric state. In the open conformation (state 1), Munc13C forms upright trimers that link the two bilayers, separating them by ∼21 nm. In the closed conformation, six copies of Munc13C interact to form a lateral hexamer elevated ∼14 nm above the bilayer. Open and closed conformations differ only by a rigid body rotation around a flexible hinge, which when performed cooperatively assembles Munc13 into a lateral hexamer (state 2) in which the key SNARE assembly-activating site of Munc13 is autoinhibited by its neighbor. We propose that each Munc13 in the lateral hexamer ultimately assembles a single SNAREpin, explaining how only and exactly six SNARE complexes are templated. We suggest that state 1 and state 2 may represent two successive states in the synaptic vesicle supply chain leading to "primed" ready-release vesicles in which SNAREpins are clamped and ready to release (state 3). | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25740.map.gz emd_25740.map.gz | 401.2 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25740-v30.xml emd-25740-v30.xml emd-25740.xml emd-25740.xml | 13.9 KB 13.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_25740.png emd_25740.png | 45.4 KB | ||
| Filedesc metadata |  emd-25740.cif.gz emd-25740.cif.gz | 6.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25740 http://ftp.pdbj.org/pub/emdb/structures/EMD-25740 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25740 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25740 | HTTPS FTP |
-Validation report
| Summary document |  emd_25740_validation.pdf.gz emd_25740_validation.pdf.gz | 287.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_25740_full_validation.pdf.gz emd_25740_full_validation.pdf.gz | 287.2 KB | Display | |
| Data in XML |  emd_25740_validation.xml.gz emd_25740_validation.xml.gz | 4.4 KB | Display | |
| Data in CIF |  emd_25740_validation.cif.gz emd_25740_validation.cif.gz | 4.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25740 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25740 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25740 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25740 | HTTPS FTP |
-Related structure data
| Related structure data |  7t7xMC  7t7cC  7t7rC  7t7vC  7t81C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25740.map.gz / Format: CCP4 / Size: 33 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25740.map.gz / Format: CCP4 / Size: 33 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Composite map created in UCSF Chimera by combining two maps from independent, focused 3D classifications using the "vop maximum" command | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
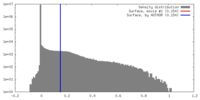
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : 2D crystal of Munc13-1 C1-C2B-MUN-C2C domains between two lipid b...
| Entire | Name: 2D crystal of Munc13-1 C1-C2B-MUN-C2C domains between two lipid bilayers. |
|---|---|
| Components |
|
-Supramolecule #1: 2D crystal of Munc13-1 C1-C2B-MUN-C2C domains between two lipid b...
| Supramolecule | Name: 2D crystal of Munc13-1 C1-C2B-MUN-C2C domains between two lipid bilayers. type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 130 KDa |
-Macromolecule #1: Protein unc-13 homolog A
| Macromolecule | Name: Protein unc-13 homolog A / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 130.895867 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GPLGSEFMAG ITSALASSTL NNEELKNHVY KKTLQALIYP ISCTTPHNFE VWTATTPTYC YECEGLLWGI ARQGMRCTEC GVKCHEKCQ DLLNADCLQR AAEKSSKHGA EDRTQNIIMV LKDRMKIRER NKPEIFELIQ EVFAVTKSAH TQQMKAVKQS V LDGTSKWS ...String: GPLGSEFMAG ITSALASSTL NNEELKNHVY KKTLQALIYP ISCTTPHNFE VWTATTPTYC YECEGLLWGI ARQGMRCTEC GVKCHEKCQ DLLNADCLQR AAEKSSKHGA EDRTQNIIMV LKDRMKIRER NKPEIFELIQ EVFAVTKSAH TQQMKAVKQS V LDGTSKWS AKISITVVCA QGLQAKDKTG SSDPYVTVQV GKTKKRTKTI YGNLNPVWEE NFHFECHNSS DRIKVRVWDE DD DIKSRVK QRFKRESDDF LGQTIIEVRT LSGEMDVWYN LDKRTDKSAV SGAIRLHISV EIKGEEKVAP YHVQYTCLHE NLF HFVTDV QNNGVVKIPD AKGDDAWKVY YDETAQEIVD EFAMRYGVES IYQAMTHFAC LSSKYMCPGV PAVMSTLLAN INAY YAHTT ASTNVSASDR FAASNFGKER FVKLLDQLHN SLRIDLSMYR NNFPASSPER LQDLKSTVDL LTSITFFRMK VQELQ SPPR ASQVVKDCVK ACLNSTYEYI FNNCHELYGR EYQTDPAKKG EVPPEEQGPS IKNLDFWSKL ITLIVSIIEE DKNSYT PCL NQFPQELNVG KISAEVMWSL FAQDMKYAME EHDKHRLCKS ADYMNLHFKV KWLYNEYVAE LPTFKDRVPE YPAWFEP FV IQWLDENEEV SRDFLHGALE RDKKDGFQQT SEHALFSCSV VDVFSQLNQS FEIIKKLECP DPQIVGHYMR RFAKTISN V LLQYADIVSK DFASYCSKEK EKVPCILMNN TQQLRVQLEK MFEAMGGKEL DAEASGTLKE LQVKLNNVLD ELSHVFATS FQPHIEECVR QMGDILSQVK GTGNVPASAC SSVAQDADNV LQPIMDLLDS NLTLFAKICE KTVLKRVLKE LWKLVMNTME RTIVLPPEF LSKLKDHMVR EEAKSLTPKQ CAVVELALDT IKQYFHAGGV GLKKTFLEKS PDLQSLRYAL SLYTQATDLL I KTFVQTQS AQGSGVEDPV GEVSVHVELF THPGTGEQKV TVKVVAANDL KWQTSGIFRP FIEVNIVGPQ LSDKKRKFAT KS KNNSWAP KYNESFQFSL SADAGPECYE LQVCVKDYCF AREDRTVGLA VLQLRELAQR GSAACWLPLG RRIHMDDTGL TVL RILSQR SNDEVAKEFV KLKSDTRSAE EGGAAPAP UniProtKB: Protein unc-13 homolog A, Protein unc-13 homolog A |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | 2D array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Component:
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 281 K / Instrument: FEI VITROBOT MARK IV / Details: blot for 5 sec before plunging, blot force -1. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 3.1 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 5.0 µm / Nominal defocus min: 3.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 10.0 Å / Resolution method: OTHER / Software - Name: RELION (ver. 3.1) Details: Combined map of best classes from two independent 3D classifications in RELION 3.1 of selected regions without angular searches using C3 symmetry expanded dataset (126711 subtomograms) and ...Details: Combined map of best classes from two independent 3D classifications in RELION 3.1 of selected regions without angular searches using C3 symmetry expanded dataset (126711 subtomograms) and separation into 8 classes. Number subtomograms used: 126711 |
|---|---|
| Extraction | Number tomograms: 62 / Number images used: 70467 Details: Particles were extracted and refined using Warp/M software |
| Final 3D classification | Number classes: 8 |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1) |
-Atomic model buiding 1
| Details | Model for fitting was generated by AlphaFold using the construct's amino acid sequence. Flexible fitting into 3D map densities was performed using ISOLDE tool in ChimeraX. |
|---|---|
| Refinement | Protocol: FLEXIBLE FIT |
| Output model |  PDB-7t7x: |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)