+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-24268 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structures of human ghrelin receptor-Gi complexes with ghrelin and a synthetic agonist | ||||||||||||
 マップデータ マップデータ | CryoEM of GHSR-Gi-MK0677 | ||||||||||||
 試料 試料 |
| ||||||||||||
 キーワード キーワード | GPCR / appetite / energy homeostasis / reward signaling / MEMBRANE PROTEIN | ||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報growth hormone secretagogue receptor activity / regulation of hindgut contraction / regulation of growth hormone secretion / negative regulation of locomotion involved in locomotory behavior / growth hormone-releasing hormone receptor activity / positive regulation of small intestinal transit / regulation of gastric motility / response to follicle-stimulating hormone / regulation of transmission of nerve impulse / ghrelin secretion ...growth hormone secretagogue receptor activity / regulation of hindgut contraction / regulation of growth hormone secretion / negative regulation of locomotion involved in locomotory behavior / growth hormone-releasing hormone receptor activity / positive regulation of small intestinal transit / regulation of gastric motility / response to follicle-stimulating hormone / regulation of transmission of nerve impulse / ghrelin secretion / growth hormone secretion / negative regulation of macrophage apoptotic process / positive regulation of small intestine smooth muscle contraction / negative regulation of norepinephrine secretion / positive regulation of eating behavior / positive regulation of appetite / adult feeding behavior / negative regulation of appetite / actin polymerization or depolymerization / positive regulation of multicellular organism growth / cellular response to thyroid hormone stimulus / positive regulation of insulin-like growth factor receptor signaling pathway / response to growth hormone / regulation of postsynapse organization / response to L-glutamate / positive regulation of vascular endothelial cell proliferation / negative regulation of interleukin-1 beta production / response to dexamethasone / positive regulation of fatty acid metabolic process / response to food / cellular response to insulin-like growth factor stimulus / regulation of synapse assembly / positive regulation of sprouting angiogenesis / regulation of neurotransmitter receptor localization to postsynaptic specialization membrane / decidualization / negative regulation of interleukin-6 production / peptide hormone binding / negative regulation of tumor necrosis factor production / postsynaptic modulation of chemical synaptic transmission / adenylate cyclase inhibitor activity / positive regulation of protein localization to cell cortex / T cell migration / positive regulation of relaxation of smooth muscle / Adenylate cyclase inhibitory pathway / hormone-mediated signaling pathway / D2 dopamine receptor binding / response to hormone / adenylate cyclase-inhibiting serotonin receptor signaling pathway / G protein-coupled serotonin receptor binding / insulin-like growth factor receptor signaling pathway / cellular response to forskolin / Peptide ligand-binding receptors / regulation of mitotic spindle organization / chemokine-mediated signaling pathway / synaptic membrane / Regulation of insulin secretion / neuropeptide signaling pathway / response to prostaglandin E / positive regulation of cholesterol biosynthetic process / negative regulation of insulin secretion / G protein-coupled receptor binding / negative regulation of inflammatory response / response to peptide hormone / G protein-coupled receptor activity / Schaffer collateral - CA1 synapse / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / G-protein beta/gamma-subunit complex binding / centriolar satellite / cellular response to insulin stimulus / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / GDP binding / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / response to estradiol / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding 類似検索 - 分子機能 | ||||||||||||
| 生物種 |  Homo sapiens (ヒト) / Homo sapiens (ヒト) /  | ||||||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.7 Å | ||||||||||||
 データ登録者 データ登録者 | Liu H / Sun D | ||||||||||||
| 資金援助 |  米国, 3件 米国, 3件
| ||||||||||||
 引用 引用 |  ジャーナル: Nat Commun / 年: 2021 ジャーナル: Nat Commun / 年: 2021タイトル: Structural basis of human ghrelin receptor signaling by ghrelin and the synthetic agonist ibutamoren. 著者: Heng Liu / Dapeng Sun / Alexander Myasnikov / Marjorie Damian / Jean-Louis Baneres / Ji Sun / Cheng Zhang /   要旨: The hunger hormone ghrelin activates the ghrelin receptor GHSR to stimulate food intake and growth hormone secretion and regulate reward signaling. Acylation of ghrelin at Ser3 is required for its ...The hunger hormone ghrelin activates the ghrelin receptor GHSR to stimulate food intake and growth hormone secretion and regulate reward signaling. Acylation of ghrelin at Ser3 is required for its agonistic action on GHSR. Synthetic agonists of GHSR are under clinical evaluation for disorders related to appetite and growth hormone dysregulation. Here, we report high-resolution cryo-EM structures of the GHSR-G signaling complex with ghrelin and the non-peptide agonist ibutamoren as an investigational new drug. Our structures together with mutagenesis data reveal the molecular basis for the binding of ghrelin and ibutamoren. Structural comparison suggests a salt bridge and an aromatic cluster near the agonist-binding pocket as important structural motifs in receptor activation. Notable structural variations of the G and GHSR coupling are observed in our cryo-EM analysis. Our results provide a framework for understanding GHSR signaling and developing new GHSR agonist drugs. | ||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_24268.map.gz emd_24268.map.gz | 118 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-24268-v30.xml emd-24268-v30.xml emd-24268.xml emd-24268.xml | 17.5 KB 17.5 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_24268.png emd_24268.png | 64.5 KB | ||
| Filedesc metadata |  emd-24268.cif.gz emd-24268.cif.gz | 6.8 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24268 http://ftp.pdbj.org/pub/emdb/structures/EMD-24268 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24268 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24268 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_24268.map.gz / 形式: CCP4 / 大きさ: 125 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_24268.map.gz / 形式: CCP4 / 大きさ: 125 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | CryoEM of GHSR-Gi-MK0677 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.826 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
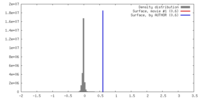
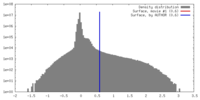
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
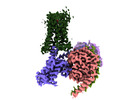
-全体 : Complex of GHSR-Gi-ghrelin
| 全体 | 名称: Complex of GHSR-Gi-ghrelin |
|---|---|
| 要素 |
|
-超分子 #1: Complex of GHSR-Gi-ghrelin
| 超分子 | 名称: Complex of GHSR-Gi-ghrelin / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1-#5 |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
-分子 #1: Guanine nucleotide-binding protein G(i) subunit alpha-1
| 分子 | 名称: Guanine nucleotide-binding protein G(i) subunit alpha-1 タイプ: protein_or_peptide / ID: 1 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 40.429059 KDa |
| 組換発現 | 生物種: Insect expression vector pBlueBacmsGCA1His (その他) |
| 配列 | 文字列: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKSTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI ...文字列: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKSTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI PTQQDVLRTR VKTTGIVETH FTFKDLHFKM FDVGGQRSER KKWIHCFEGV TAIIFCVALS DYDLVLAEDE EM NRMHESM KLFDSICNNK WFTDTSIILF LNKKDLFEEK IKKSPLTICY PEYAGSNTYE EAAAYIQCQF EDLNKRKDTK EIY THFTCA TETKNVQFVF DAVTDVIIKN NLKDCGLF UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-1 |
-分子 #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| 分子 | 名称: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 タイプ: protein_or_peptide / ID: 2 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 37.417918 KDa |
| 組換発現 | 生物種: Insect expression vector pBlueBacmsGCA1His (その他) |
| 配列 | 文字列: MSELDELRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR QGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL ...文字列: MSELDELRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR QGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL WDIETGQQTT TFTGHTGDVM SLSLAPDTRL FVSGACDASA KLWDVREGMC RQTFTGHESD INAICFFPDG NA FATGSDD ATCRLFDLRA DQELMTYSHD NIICGITSVS FSKSGRLLLA GYDDFNCNVW DALKADRAGV LAGHDNRVSC LGV TDDGMA VATGSWDSFL KIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-分子 #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| 分子 | 名称: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 タイプ: protein_or_peptide / ID: 3 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 7.859173 KDa |
| 組換発現 | 生物種: Insect expression vector pBlueBacmsGCA1His (その他) |
| 配列 | 文字列: MASNNTASIA QARKLVQQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASQNP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-分子 #4: Antibody fragment
| 分子 | 名称: Antibody fragment / タイプ: protein_or_peptide / ID: 4 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 26.236244 KDa |
| 組換発現 | 生物種: Insect expression vector pBlueBacmsGCA1His (その他) |
| 配列 | 文字列: VQLVESGGGL VQPGGSRKLS CSASGFAFSS FGMHWVRQAP EKGLEWVAYI SSGSGTIYYA DTVKGRFTIS RDDPKNTLFL QMTSLRSED TAMYYCVRSI YYYGSSPFDF WGQGTTLTVS SGGGGSGGGG SGGGGSDIVM TQATSSVPVT PGESVSISCR S SKSLLHSN ...文字列: VQLVESGGGL VQPGGSRKLS CSASGFAFSS FGMHWVRQAP EKGLEWVAYI SSGSGTIYYA DTVKGRFTIS RDDPKNTLFL QMTSLRSED TAMYYCVRSI YYYGSSPFDF WGQGTTLTVS SGGGGSGGGG SGGGGSDIVM TQATSSVPVT PGESVSISCR S SKSLLHSN GNTYLYWFLQ RPGQSPQLLI YRMSNLASGV PERFSGSGSG TAFTLTISRL EAEDVGVYYC MQHLEYPLTF GA GTKLEL |
-分子 #5: Growth hormone secretagogue receptor type 1
| 分子 | 名称: Growth hormone secretagogue receptor type 1 / タイプ: protein_or_peptide / ID: 5 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 41.392383 KDa |
| 組換発現 | 生物種: Insect expression vector pBlueBacmsGCA1His (その他) |
| 配列 | 文字列: MWNATPSEEP GFNLTLADLD WDASPGNDSL GDELLQLFPA PLLAGVTATC VALFVVGIAG NLLTMLVVSR FRELRTTTNL YLSSMAFSD LLIFLCMPLD LVRLWQYRPW NFGDLLCKLF QFVSESCTYA KVLTITALSV ERYFAICFPL RAKVVVTKGR V KLVIFVIW ...文字列: MWNATPSEEP GFNLTLADLD WDASPGNDSL GDELLQLFPA PLLAGVTATC VALFVVGIAG NLLTMLVVSR FRELRTTTNL YLSSMAFSD LLIFLCMPLD LVRLWQYRPW NFGDLLCKLF QFVSESCTYA KVLTITALSV ERYFAICFPL RAKVVVTKGR V KLVIFVIW AVAFCSAGPI FVLVGVEHEN GTDPWDTNEC RPTEFAVRSG LLTVMVWVSS IFFFLPVFCL TVLYSLIGRK LW RRRRGDA VVGASLRDQN HKQTVKMLAV VVFAFILCWL PFHVGRYLFS KSFEPGSLEI AQISQYCNLV SFVLFYLSAA INP ILYNIM SKKYRVAVFR LLGFEPFSQR KLSTLKDESS RAWTESSINT UniProtKB: Growth hormone secretagogue receptor type 1 |
-分子 #6: 1-(methanesulfonyl)-1'-(2-methyl-L-alanyl-O-benzyl-D-seryl)-1,2-d...
| 分子 | 名称: 1-(methanesulfonyl)-1'-(2-methyl-L-alanyl-O-benzyl-D-seryl)-1,2-dihydrospiro[indole-3,4'-piperidine] タイプ: ligand / ID: 6 / コピー数: 1 / 式: 1KD |
|---|---|
| 分子量 | 理論値: 528.664 Da |
| Chemical component information | 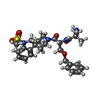 ChemComp-1KD: |
-分子 #7: CHOLESTEROL
| 分子 | 名称: CHOLESTEROL / タイプ: ligand / ID: 7 / コピー数: 1 / 式: CLR |
|---|---|
| 分子量 | 理論値: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 平均電子線量: 82.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー














































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)